Demand for genetically improved lines is strong in Southeast Asia

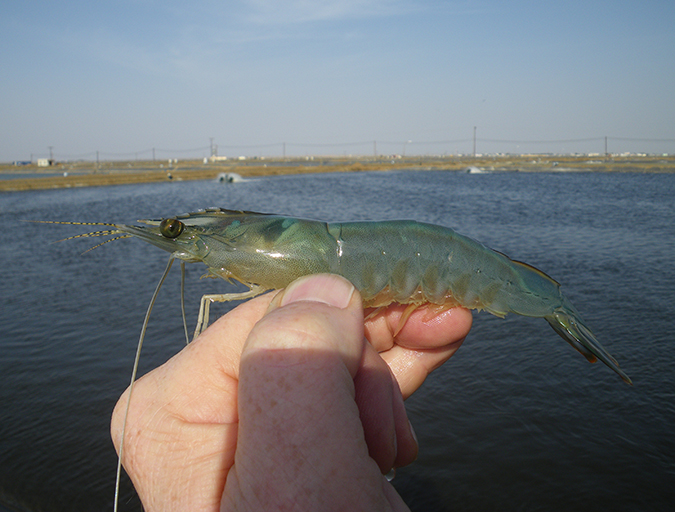
Australian shrimp farms primarily grow black tiger shrimp, Penaeus monodon, for which elite genetic lines have been successfully produced by selective breeding. As reported by Brett Glencross and co-authors in Aquaculture Nutrition in 2013, these elite lines provide harvest yields more than double that of unselected lines and have improved survival and growth performance, and lowered metabolic rates and energy requirements.
The demand to access these genetically improved lines is strong from the countries of Southeast Asia. However, the Australian shrimp-farming industry is highly motivated to prevent unlicensed breeding. As there are currently no mechanisms to confer failproof reproductive sterility and thus genetic copyright on a commercial scale in penaeid shrimp, this has precluded the sale of elite stocks beyond the individual farm enterprises that breed them. Remarkably little is known about the underlying biochemical and genetic processes that control fertility, sex and germ line determination, or other aspects of shrimp development.
To develop an alternative mechanism for genetic copyright, the authors used a bioinformatics approach to identify germ line genes that could be potentially targeted to ablate the germ line. This approach has also resulted in the production of developmental transcriptomes for penaeid shrimp.
Preparing embryos for genetic sequencing
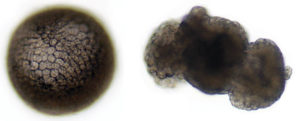
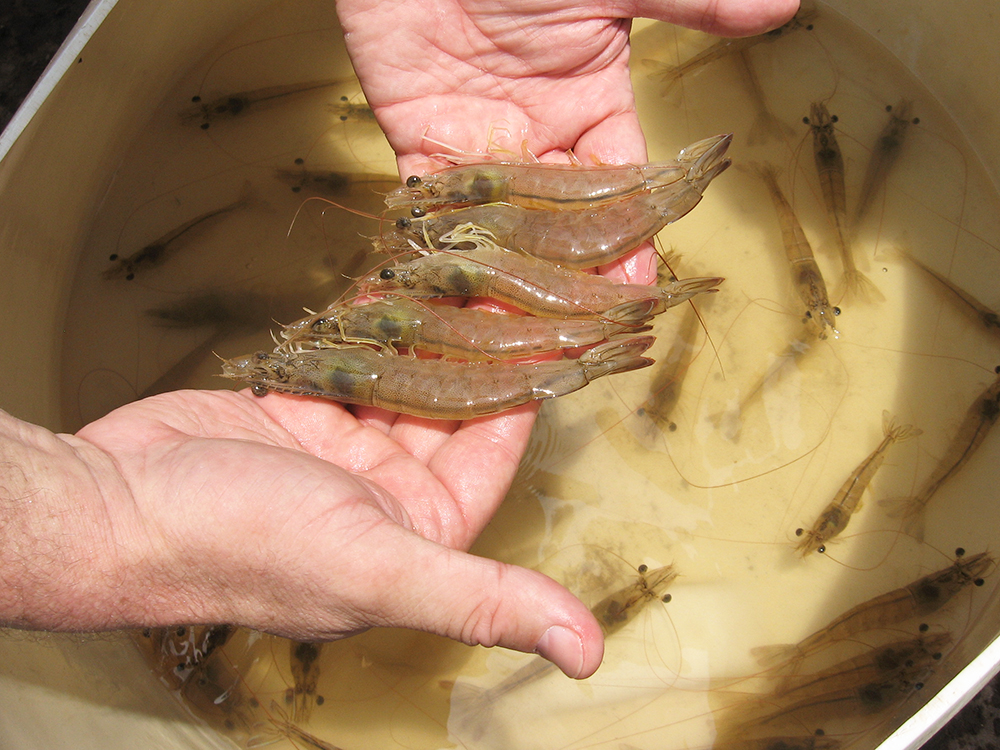
The kuruma shrimp, Penaeus japonicas, is readily spawned at one of the research stations of Australia’s Commonwealth Science and Industrial Research Organisation, so this species was used as a model. It was first demonstrated by Takao Kajishima in 1951 that when P. japonicus embryos were separated by a sharp needle at the two-cell stage, the “animal” half only developed into a hollow ball of cells, while the “vegetal” half continued developing.
By utilizing this technique, researchers had the opportunity to look for embryonic genes differentially expressed in animal or vegetal half-embryos. They predicted that germ line genes might be enriched in the vegetal half, since the primordial germ cell was hypothesized to arise from vegetal determinants.
P. japonicus embryos were separated at the two-cell stage and allowed to develop until the animal and vegetal half-embryos could be distinguished. The animal and vegetal half-embryos were pooled separately, total ribonucleic acid was isolated and reverse transcribed to complementary DNA, and the resulting transcriptomes were sequenced.
Reads from each library were assembled, annotated and screened for known germ line and mesoderm genes. Pre-existing P. monodon ovary and nauplius transcriptome libraries were also screened for developmental genes of interest.
Germ line genes
The germ line genes vasa and nanos were previously found in shrimp by cloning methods, but neither vasa nor nanos appeared in the embryo transcriptomes. The authors found other candidate germ line genes, including pumilio, germ cell-less, staufen and tudor, in both animal and vegetal transcriptomes. All four of these were more highly expressed in ovary and/or testes than in other adult tissues and could be detected during embryonic development. Next, the authors looked for where the selected germ line genes were expressed in the developing embryos.
Custom monoclonal antibodies were generated to examine the protein expression of shrimp vasa, nanos, pumilio, and germ cell-less genes by immunoblotting and immunolocalization. The vasa and nanos antibodies labeled a structure in embryos previously hypothesized to be a germ granule. This structure has been detected is several shrimp species and is inherited by embryonic cells hypothesized to give rise to the primordial germ cell. The pumilio and germ cell-less antibodies are still being characterized.
Mesoderm genes
The mesoderm genes twist, snail, mef-2 and brachyury were found in the P. japonicus half-embryo transcriptomes. These all function as transcription factors, which activate or repress other genes in mesoderm, muscle or related tissues. The twist and brachyury genes were found only in the vegetal transcriptomes, while snail and mef-2 were found in both animal and vegetal transcriptomes.
Recently, five developmental transcriptomes – embryo, nauplius, zoea, mysis and postlarva – from the Pacific white shrimp, Litopenaeus vannamei, were published in the gene databases by Jianhai Xiang’s laboratory. The L. vannamei versions of these mesoderm genes were identified, and primers were developed in collaboration with the Xiang lab to study their expression by quantitative polymerase chain reaction in this species.
The expression of the mesoderm genes is consistent with their known functions as transcription factors in other organisms. For twist and snail, expression was not detected until later embryonic stages and continued into the larval and postlarval stages. The mef-2 expression was detected at all stages of development from zygote to postlarva. For brachyury, expression was highest during gastrulation – consistent with its known role in promoting cell movements during embryogenesis.
The next step will be to study the expression patterns of these genes in embryos and larvae, and antibodies to shrimp twist, mef2 and brachyury are in production. In the model amphipod crustacean Parhyale hawaiensis, work by Nipam Patel’s laboratory has shown that twist and snail proteins are expressed in the posterior mesoderm, while mef-2 protein is expressed in posterior mesoderm and persists in developing muscles.
Application: germ line knockdown
The availability of germ line gene sequences and antibodies to detect their protein products allows for experiments to ablate the shrimp germ line by targeted gene knockdown.
RNA interference is a powerful and highly accurate natural biological pathway in shrimp that can be utilized for such studies. This can be performed by administration of double-stranded RNA via tail-muscle injection or oral delivery.
The mesoderm/muscle gene sequences and molecular tools developed in parallel should be useful in further understanding the basic biology of mesoderm and muscle development in shrimp, for example in detecting cellular changes in faster-growing genetic lines.
(Editor’s Note: This article was originally published in the July/August 2015 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Philip L. Hertzler, Ph.D.
Professor
Central Michigan University
Department of Biology
Brooks Hall 217
Mount Pleasant, Michigan 48858 USA[117,100,101,46,104,99,105,109,99,64,108,112,49,122,116,114,101,104]
-
Melony J. Sellars, Ph.D.
Senior Research Scientist, Research Manager
ARC Research Hub for Advanced Prawn Breeding
Integrated Sustainable Aquaculture Production
Agriculture Flagship, CSIRO EcoSciences Precinct
Brisbane, Queensland, Australia
Tagged With
Related Posts

Health & Welfare
New management tools for EHP in penaeid shrimp
Authors examined the histological features from shrimp infected with the emerging microsporidian parasite Enterocytozoon hepatopenaei (EHP). A PCR assay method was used to detected in hepatopancreatic tissue, feces and water sampled from infected shrimp tanks, and in some samples of Artemia biomass.

Aquafeeds
Biofloc consumption by Pacific white shrimp postlarvae
The stable isotopes technique with δ13C and δ15N can be used to determine the relevance of different food sources to shrimp feeding during the pre-nursery phase of Litopenaeus vannamei culture. During this trial, different types of commercial feed, microalgae, Artemia sp. nauplii and bioflocs were used as food sources.

Health & Welfare
Saudi Arabia developing effective farmed shrimp biosecurity strategy
Biosecurity strategies adopted by the shrimp industry in the Kingdom of Saudi Arabia have resulted in historical production in 2015 and a similar forecast for 2016, and are a significant step towards the long-term goal of sustainable aquaculture.

Health & Welfare
Non-invasive diagnostic tool developed for shrimp disease EMS
The presence of AHPND-Vibrio parahaemolyticus can be detected both in fecal DNA samples and in the enriched bacterial broth with samples of enrichment broth showing increased sensitivity.



