Authentication of fisheries products based on distinct methods of DNA extraction

The development of modern tools that assure the identity of seafood sold for human consumption is an important area of applied research to help prevent the trade of mislabeled products. Several studies have shown the relevance of genetic science in detecting substitutions and investigating the authenticity of processed products, demonstrating that molecular tools can assist in inspection practices, provide DNA-based certification of the marketed product and ensure monitoring of the fishing industry.
An essential step in genetic and molecular biology studies is obtaining high-quality genetic material. Several procedures for DNA extraction samples have been developed and widely used in the forensics and identification of biological products, and these methods became particularly popular when specialized companies started producing commercial kits for DNA extraction. However, a single method is not compatible with all types of samples (biological tissue). Therefore, the selection of DNA extraction method should rely on the sample origin, the purpose of the analysis and the final targeted product.
This article – summarized from the original publication (Lutz, I. et al. 2023. Quality analysis of genomic DNA and authentication of fisheries products based on distinct methods of DNA extraction. PLoS ONE 18(2): e0282369) – presents the results of a study that compared three different protocols of DNA extraction: NaCl (saline); phenol-chloroform and a commercial kit (Promega) from three biological tissues of five individuals of Caribbean red snapper (Lutjanus purpureus) under two methods of storage.
Study setup
This study was conducted at the Laboratório de Genética Aplicada (LAGA), Instituto de Estudos Costeiros (IECOS), at Universidade Federal do Pará (UFPA)/Bragança, Brazil. The species selected for the study was the Caribbean red snapper (L. purpureus), one of the most exploited fish species from the northern coast of Brazil and highly commercialized as fillets or frozen specimens for national and international trade. The samples comprised five individuals of red snapper obtained from a fisheries port in Bragança (the northeastern region of the state of Pará, Brazil). This region is Brazil’s leading trade center of red snappers, providing processed fisheries products for several markets worldwide, such as the Caribbean, USA, and Europe.
The tissue samples collected for analysis included pieces of muscle, tongue, and caudal fins stored at frozen conditions (alcohol-free) or in 90 percent ethanol to be tested. Three commonly used protocols of DNA extraction from three biological tissues conserved at distinct conditions were compared: (1) extraction using saline solution (NaCl); (2) the phenol-chloroform-isoamyl alcohol method; and (3) the commercial Wizard® Genomic kit (Promega).
A schematic representation of the experimental design is presented in Fig. 1, showing the main steps for the accomplishment of this work, from the selection of the species and removal of the biological tissues to the observation of the different variables tested for each extraction protocol. We extracted three different types of tissue and used two forms of storage (Fig. 1A) to observe a possible variation in the quality of the extracted DNA. In order to standardize the DNA extraction protocol for the molecular certification chain of processed fish, three methods were compared (Fig. 1B), as well as the cost, time to process the samples, quality and quantity of extracted DNA (Fig. 1C) and generation of functional sequences with the DNA barcode tool (Fig. 1D).
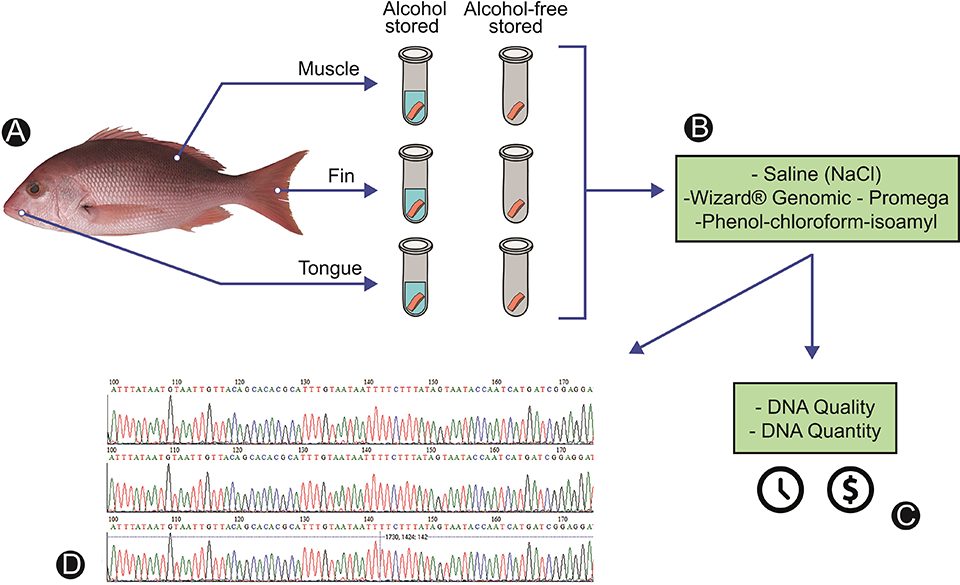
For detailed information on the experimental design, sample collection and preservation; DNA extraction with the three methods evaluated; DNA analyses; and statistical analyses, consult the original publication.
Results and discussion
The amount and quality (integrity and purity) of extracted DNA are critical factors to their successful application and reproducibility in molecular genetics. The L. purpureus we analyzed were acquired directly at the time of landing from a commercial fleet and we did not obtain information on the time elapsed between capture and collection of tissues for storage, which may influence DNA degradation. We recommend this information is collected in future studies when available. The method based on saline solution and the commercial Wizard® Genomic kit (Promega) yielded the highest integrity (Fig. 2) and the highest concentration values in the DNA samples.
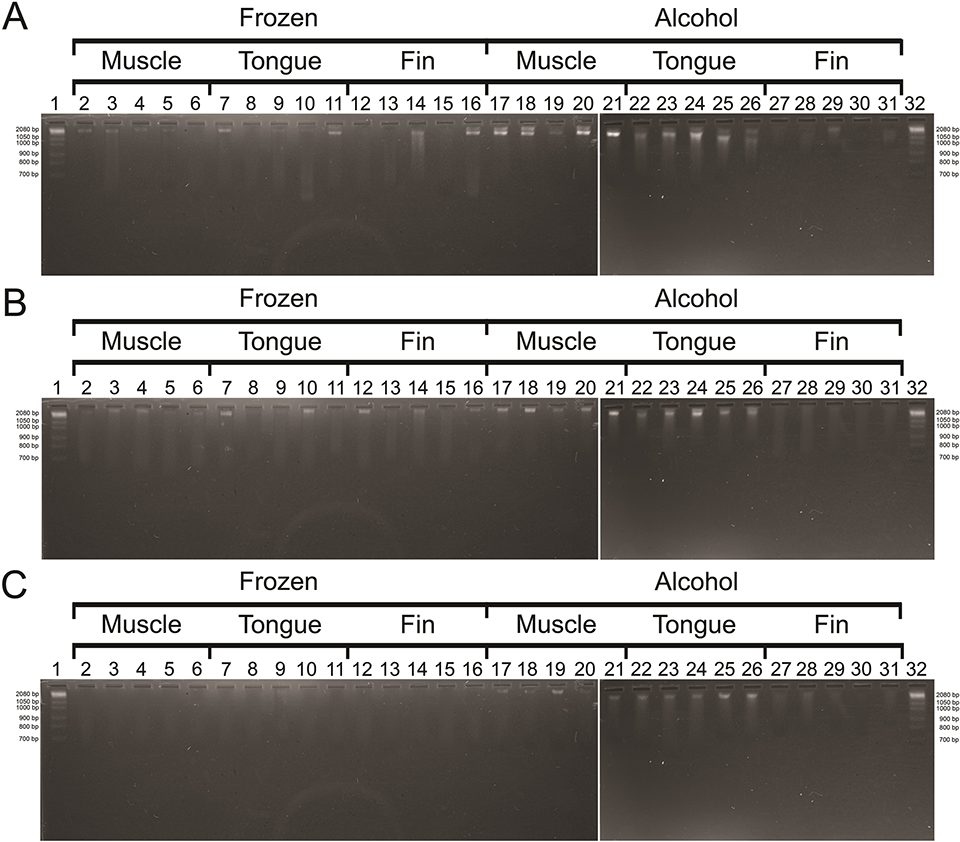
Based on the visualization in agarose gel, the best DNA bands profile considering their integrity and high molecular weight was observed in the protocols based on saline and the commercial kit despite some variation according to tissue and storage condition (Figs. 2A and 2B). On the other hand, the phenol-chloroform method yielded many samples with low DNA concentrations (Fig. 2C).
Concerning the storage conditions, the best results were observed in samples preserved in alcohol rather than just frozen using the three protocols of DNA extraction. By preserving tissues in alcohol, bacterial proliferation and the degradation of nucleic acids by the activity of various enzymes are avoided. The preparation of tissues stored in alcohol increases the chances of obtaining the best results.
We observed the highest mean values in DNA concentration in tongue samples for the protocols of DNA extraction based on saline and the commercial kit and fins for the phenol-chloroform method. The use of tongue and fins for DNA extraction preserves the external morphology of fishes, thus avoiding depreciation in trade markets without compromising their certification, representing a significant added value, particularly in exported products. In addition, using fin or tongue tissues is also recommended to obtain DNA samples from museum specimens that cannot be damaged.
The purity of DNA samples extracted using the three methods of DNA extraction in this study was variable. The saline method and the Wizard® Genomic kit (Promega) yielded satisfactory DNA purity levels, with the saline method showing to be the most suitable for DNA extraction.
New circular DNA vaccines and heritable, anti-viral immunity possible for shrimp
The development of commercial kits made it possible to shorten the time spent in DNA extraction compared to other published protocols, such as those based on saline solution and phenol-chloroform. Most DNA extraction kits require ready-to-use reagents, reducing the time in preparing the material and demanding a low number of operational steps, making the whole process simple and easily performed even by an untrained technician. On the other hand, while commercial kits are time-saving and straightforward methods, their utilization increases the cost associated with the extraction of an individual sample, representing an increase of 45.6 and 42.5 percent for the saline or the phenol-chloroform methods, respectively.
The results of the saline method yielded similar to those obtained using the Wizard® Genomic kit (Promega) concerning the quantity and quality of extracted DNA samples, besides being the least expensive method among the three methods tested. Therefore, fisheries industries with limited infrastructure or reduced budgets could rely on methods based on saline solutions to obtain high-quality DNA samples while achieving significant financial savings.
The sequences used in this pilot assay resulted in the correct identification of sampled species and the reliable discrimination of Lutjanidae in the Neighbor-Joining tree, regardless of the tested extraction method (Fig. 3), confirming their utilization as efficient authentication tools.
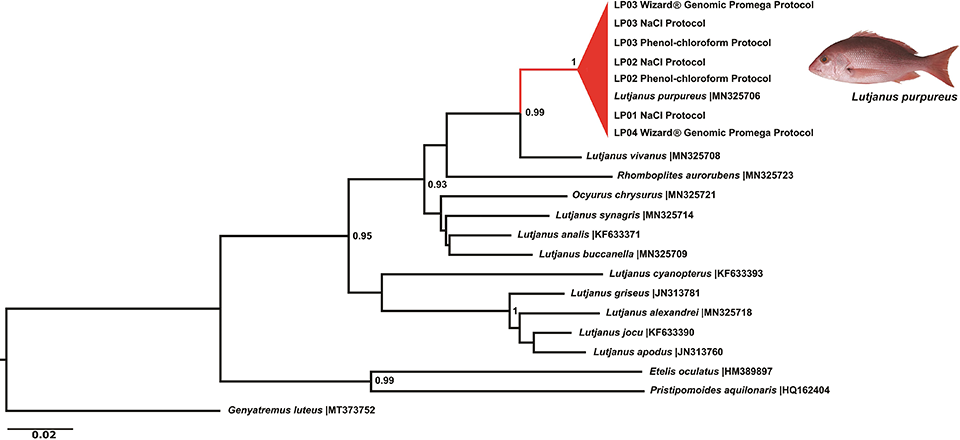
The use of molecular genetics as a tool in forensics and for the certification of commercial products has proven to be highly efficient in solving cases of seafood product substitutions. A certified product guarantees transparency for consumers, as well as food safety and traceability of exploited species, being helpful to the effective management of fisheries resources. Compared to other biotechnology tools for fish identification and traceability, the procedures based on molecular genetics still present some drawbacks, such as high costs and the time spent analyzing the samples.
However, recent advances in genetic analyses have optimized this process, reducing both the financial costs and the highly specialized labor required. Therefore, DNA-based methods represent a highly recommended tool for inspection and certification analysis, since they are more selective, sensitive and reliable than other methodologies, thus being particularly useful to the fisheries industry.
Our DNA amplification and sequencing success demonstrates that the three DNA extraction methods can be used in seafood products. However, other variables must be considered, such as time and costs, the availability of biological tissues, infrastructure, and possible toxic effects related to each method. After ensuring good yield and quality DNA in the initial extraction step, the molecular techniques steps that follow should also be standardized and optimized for better utilization by the routine fishing industry.
Perspectives
We compared three DNA extraction methods commonly used in scientific reports and by regulatory agencies in several countries. Both the saline protocol and commercial kit provided similar results concerning the quantity and quality of extracted DNA. Therefore, the final choice would depend on the available financial resources and time for carrying out each procedure. While the commercial kit requires less time to isolate DNA samples, the costs of reagents increase considerably. On the other hand, the protocol based on the saline solution is less expensive even though it requires a relatively long period to be performed, being suitable for small fisheries industries.
Concerning the biological tissue and storage form, we propose tongue and fin, and storage in alcohol as indicated for best results in concentration and purity of extracted DNA. Finally, we recommend that new comparisons between extraction protocols be performed in future studies, inserting new commercial kits and maintaining the published classical protocols. These protocols should also be tested on other fishery products, such as cephalopods, mollusks and crustaceans, as well as on tissues stored in other conditions.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Grazielle Evangelista Gomes, Ph.D.
Corresponding author
Laboratório de Genética Aplicada, Instituto de Estudos Costeiros, Universidade Federal do Pará, Bragança, Pará, Brasil[114,98,46,97,112,102,117,64,101,108,108,101,105,122,97,114,103]
Tagged With
Related Posts

Health & Welfare
Effects of dietary lipid on the gut microbiota of first feeding Atlantic salmon
An evaluation of dietary oils showed the occurrence of a core microbiota in early life stage of Atlantic salmon, independent of oil type in the feed.

Intelligence
Bahamas venture focuses on grouper, other high-value marine fish
A new venture under development in the Bahamas will capitalize on Tropic Seafood’s established logistics and infrastructure to diversify its operations from processing and selling wild fisheries products to include the culture of grouper and other marine fish.

Health & Welfare
Mesocosm systems for semi-intensive larval rearing of marine fish
In marine fish hatcheries, a mesocosm is a tank or pond in which a natural phytoplankton or zooplankton bloom is developed prior to stocking yolk-sac larvae.

Innovation & Investment
AquaGen CEO: Genomics are transforming aquaculture
The CEO of AquaGen knew that the Norwegian research group’s work in genomics was key to the salmon industry’s future. And that was before she even worked there.


![Ad for [BSP]](https://www.globalseafood.org/wp-content/uploads/2025/07/BSP_B2B_2025_1050x125.jpg)
