Valuable insights for developing health-oriented and sustainable aquaculture strategies for P. vannamei in high-salinity environments

How do extreme salinity gradients shape the intestinal microbiota of Pacific white shrimp (Penaeus vannamei)? A shrimp’s gut microbiota is essential for supporting host health and enabling adaptation to environmental stresses. However, how the intestinal microbial community in P. vannamei responds to elevated salinity levels is still not well characterized.
In China, Xiangli Tian and co-workers employed metagenomic sequencing – a technique that directly sequences and analyzes the collective DNA from all microorganisms present in a complex environmental or biological sample without the need to culture or isolate individual species, allowing comprehensive study of microbial community structure, diversity and functional potential – to examine and compare the structural composition and functional capabilities of the gut bacteria in shrimp maintained under low (L-), medium (M-), and high (H-) salinity conditions.
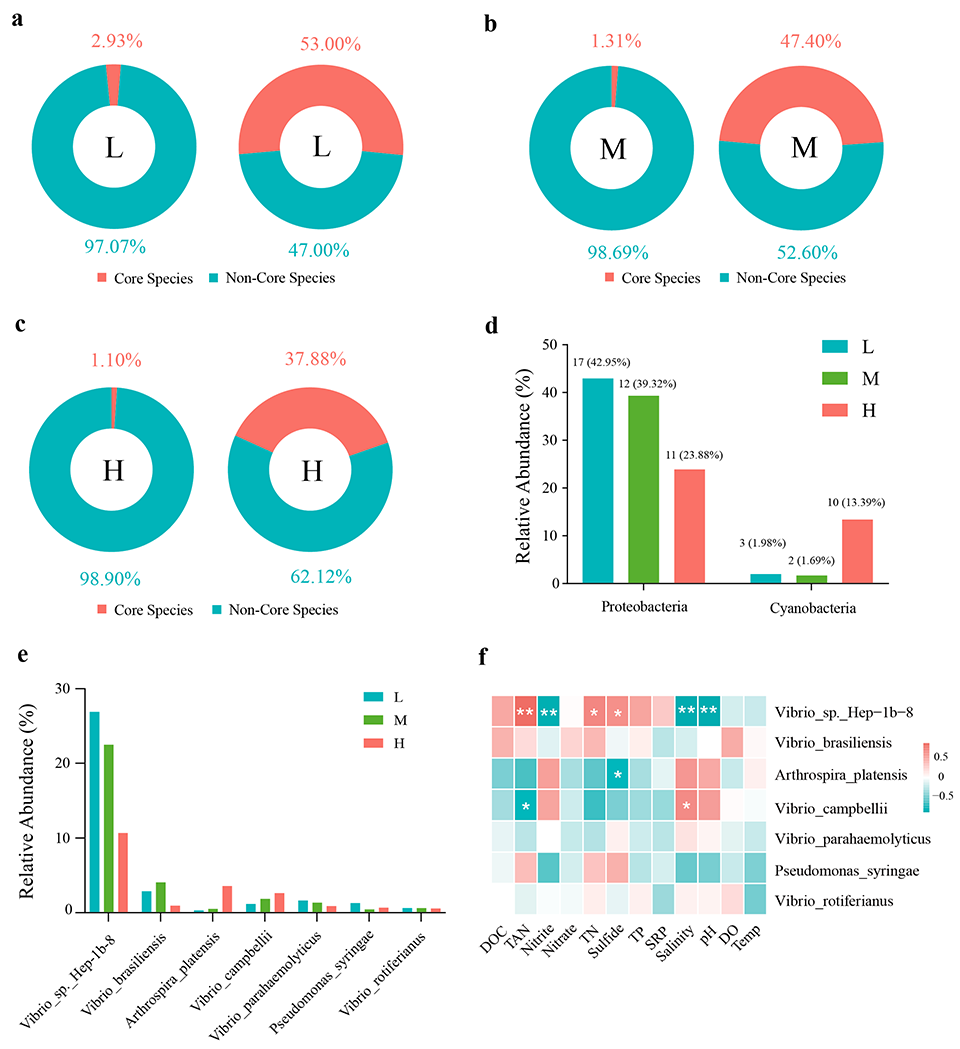
Alpha diversity – a measure of the richness and evenness of species or taxa within a single sample or community – of the microbiota increased significantly as salinity rose, and principal coordinate analysis (PCoA) demonstrated distinct clustering and clear separation of the microbial communities across the three salinity groups. Analysis of core microbiota identified seven taxa shared among all groups, with five of them belonging to the genus Vibrio. Microbial source tracking analysis showed that the contribution of bacteria originating from the surrounding environment grew progressively larger with increasing salinity.
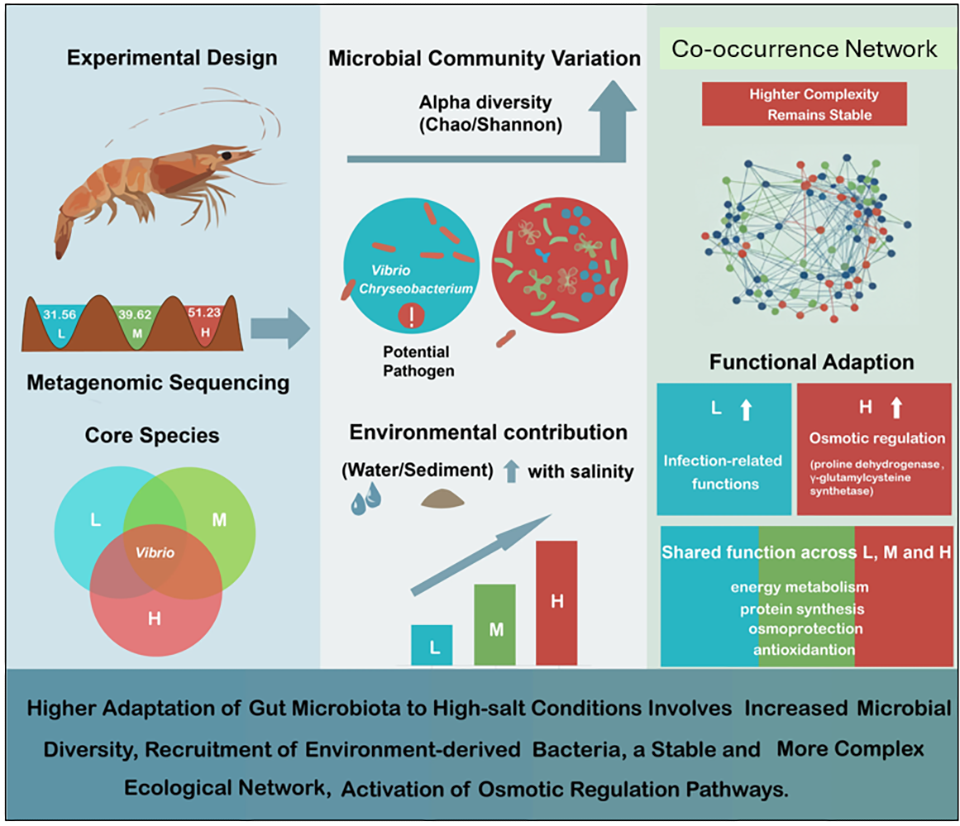
Data analysis indicated that networks in the M- and H-salinity groups were more complex in structure yet exhibited stability levels similar to those observed in the L-salinity group, the authors discovered. Notably, the low-salinity (L) group was enriched in potential pathogenic taxa (such as Vibrio and Chryseobacterium) as well as functions linked to infection and pathogenicity.
Functional profiling revealed that the high-salinity (H) group was particularly enriched in key enzymes, including proline dehydrogenase, glutamate-cysteine ligase and methyltransferases. These enzymes are interconnected in pathways involving compatible solutes which collectively play a critical role in bolstering microbial osmoprotection under hypersaline stress. And across all salinity levels, certain core functions were consistently present and related to energy metabolism, protein synthesis, osmoprotection and antioxidant defense mechanisms.
Taken together, the authors’ results provide the first simultaneous demonstration – through both structural and functional perspectives – of the potential pathogenic features dominating the intestinal microbiota under low-salinity conditions and the adaptive strategies employed by the microbiota in hypersaline (H) environments. Their insights offer valuable guidance for improving health management and disease prevention strategies in high-salinity shrimp aquaculture systems.
Relevance of research findings to the industry
Annual production of P. vannamei in hypersaline pond systems – such as those in China’s Bohai Bay and elsewhere – yields are typically 20 to 40 percent lower than in optimal conditions, largely due to salinity-induced dysbiosis (an imbalance or disruption in the composition, diversity, or function of a microbial community, such as the gut microbiota that deviates from a healthy, stable state, often leading to negative impacts on host health, immunity, or disease susceptibility) of the gut microbiota and associated disease outbreaks like acute hepatopancreatic necrosis disease (AHPND), caused by a Vibrio species.
The present study’s finding of pathogen enrichment (including species such as V. cholerae) in low-salinity environments provides actionable insights for strengthening biosecurity measures. For example, routine monitoring of total ammonia nitrogen (TAN) levels could help mitigate these risks, with potential to decrease production losses by 15–30 percent, consistent with outcomes observed in probiotic intervention trials.
On the other hand, the osmoprotective adaptations observed in the high-salinity (H) group – particularly upregulation of pathways involving various enzymes – support the feasibility of expanding shrimp farming into saline-alkaline wastelands and other marginal lands. This approach could substantially reduce reliance on freshwater resources, especially important given ongoing climate-driven salinization and the projected annual growth rate for the sector in the near future.
Practical industry applications include the development of betaine-supplemented feeds that emulate these microbial osmoprotective mechanisms, thereby improving survival rates under stressful conditions, as demonstrated in other farmed penaeid shrimp. Additionally, tailored probiotic consortia incorporating strains with functional similarities to Bacillus glennii could help maintain stable core microbiota communities, increasingly critical as Vibrio infections intensify in response to rising water temperatures.
For shrimp farming operations in northern China and elsewhere, these findings facilitate greater intensification on marginal saline lands while enabling integration of multi-omics technologies for real-time monitoring of shrimp health and early intervention.
Perspectives
This study systematically characterized the structural and functional features of the intestinal bacterial community of P. vannamei under hypersaline aquaculture conditions, revealing significant differences across salinity gradients. Bacterial alpha-diversity increased with salinity, accompanied by distinct shifts in community composition and functional profiles. Proteobacteria was the dominant phylum, with Vibrio sp. Hep-1b-8 and V. brasiliensis as the predominant species. The bacterial functions shared across salinities were primarily associated with energy metabolism, protein synthesis, osmoprotection, and antioxidant defense.
Overall, these findings advance the understanding of how shrimp intestinal bacterial communities adapt structurally and functionally to hypersaline conditions, providing valuable insights for developing health-oriented and sustainable aquaculture strategies for P. vannamei in high-salinity environments.
Fermentation technology in aquafeeds: Addressing performance through microbial innovation

Microbial fermentation can improve how aquafeeds perform for farmed fish and some crustaceans, offering potential to cut production costs and improve operational performance, while addressing challenges like disease resistance.
A review by Caihuan Ke and co-workers synthesizes the role of fermentation, which involves using microbes like lactic acid bacteria, Bacillus, yeasts and molds to process plant-based (e.g., soybean meal, cottonseed meal), animal-based (e.g., meat and bone meal, insect proteins) and energy feeds (e.g., corn, wheat bran) under controlled conditions.
Key processes include aerobic and anaerobic fermentation, with optimal parameters such as strains for efficient degradation, temperatures of 30–40 degrees-C, durations of 2–10 days, moisture levels of 60–80 percent and monitoring via crude protein changes and microbial titers.
Fermented feed represents an innovative form of functional feed that improves both the nutritional quality and palatability of aquafeeds through controlled microbial fermentation processes. It effectively addresses key challenges in aquaculture, such as reduced digestive capacity, weakened immune function in cultured species and excessive dependence on fishmeal-based formulations.
Authors examined fermented feed, its associated advantages, the main types currently in use, and the key factors that determine its effectiveness. And they also compiled evidence of its positive impacts on aquaculture species, including enhanced growth performance, strengthened immunity, favorable modulation of the intestinal microbiota and improved end-product quality.
However, although fermented feed holds substantial promise for lowering production costs and increasing overall efficiency, its adoption is currently constrained by limited large-scale industrial manufacturing and a relative scarcity of research focused on crustacean species.
To support wider implementation in aquaculture, future studies should prioritize the creation of comprehensive databases on fermentation parameters, the advancement of intelligent and automated manufacturing technologies and thorough economic assessments that evaluate cost-benefit ratios and scalability.
Relevance of research findings to the industry
The findings are highly relevant to the aquaculture industry, which faces pressures from rising fishmeal costs, sustainability demands and disease outbreaks. Fermented feeds offer a viable alternative by reducing reliance on fishmeal – substitutions of 10–100 percent have shown maintained or improved growth in species like largemouth bass, groupers and others – potentially cutting production costs by enhancing feed efficiency and profitability. Immune and gut health improvements translate to lower mortality and antibiotic use, aligning with global regulations on antimicrobial resistance. For example, fermented moringa leaves boost antioxidants, reducing oxidative stress in high-density farming.
Environmentally, fermentation degrades mycotoxins and utilizes agricultural byproducts like cassava residue, promoting circular economies and reducing waste. Product quality enhancements, such as increased iodine in salmon and better flavor profiles, could improve market value and consumer appeal.
However, industry adoption is hindered by limited data on crustaceans (e.g., shrimp, crabs), for which benefits like serine/aspartate increases (+130 percent / 30 percent) are promising but understudied. Overall, these findings support scalable, eco-friendly feed strategies, but require economic analyses to justify investments in fermentation infrastructure.
Moringa leaf extract can boost Pacific white shrimp immune responses
Perspectives
Fermented feeds improve feed utilization efficiency, immune performance, gut health and the overall quality of aquaculture products by leveraging microbial pre-digestion and probiotic effects. Fermented feeds have significant promise for lowering production costs and enhancing operational efficiency while effectively tackling common issues in farmed aquatic species, such as impaired digestion and reduced resistance to disease.
Nevertheless, broader adoption and further development of fermented feeds continue to be limited by several key obstacles, including difficulties in controlling unwanted contaminating bacteria, insufficient technical knowledge among practitioners and incomplete understanding of the precise biological mechanisms involved.
The authors concluded that to overcome these barriers and unlock the full potential, future efforts should prioritize the following: development and optimization of mixed-strain fermentation approaches, implementation of intelligent and automated production systems, expanded research focused on crustacean species (including shrimp and crabs) and strategic synergistic combinations of diverse raw materials.
These advancements will drive fermented feed toward higher efficiency, improved environmental sustainability and greater industrial scalability – ultimately delivering essential support for the long-term sustainable growth of the aquaculture sector.
A comprehensive comparative analysis of genomic prediction models in four aquaculture species

Genomic selection – a modern breeding method that uses genome-wide genetic markers to predict the breeding values or phenotypic performance of individuals for complex traits – could have major impacts in aquaculture breeding protocols.
New research by Hailiang Song and colleagues addressed the scarcity of cross-species, standardized comparisons in genomic selection, which uses genome-wide markers to predict breeding values early, accelerating genetic gains for key traits like growth and disease resistance in species with high fecundity and long generation intervals.
Genomic prediction is now widely used in selective breeding programs for aquaculture species, yet comprehensive, systematic comparisons of prediction accuracy across multiple species and diverse analytical methods – conducted within a single, standardized framework – are still scarce.
This study performed a thorough cross-species evaluation of genomic prediction performance in four major aquaculture species: Atlantic salmon (Salmo salar), gilthead sea bream (Sparus aurata), common carp (Cyprinus carpio) and rainbow trout (Oncorhynchus mykiss). Ten different genomic prediction models were tested, encompassing Genomic Best Linear Unbiased Prediction (GBLUP; a widely used genomic prediction method in animal and plant breeding that estimates breeding values using a genomic relationship matrix instead of a pedigree-based relationship matrix), several Bayesian approaches (a family of statistical modeling methods that incorporate prior knowledge or beliefs about parameters and update them with observed data, allowing flexible handling of uncertainty, variable shrinkage, and marker effect heterogeneity), and machine learning techniques.
Prediction accuracies varied substantially across species and models, ranging from 0.49 to 0.85, and showed a strong positive correlation with trait heritability. Traits with higher heritability consistently yielded superior prediction accuracies: Rainbow trout and common carp achieved the highest overall performance (0.75–0.83 and 0.73–0.85, respectively), while Atlantic salmon and gilthead sea bream displayed lower and more variable accuracies (0.49–0.61 and 0.49–0.66).
No individual model proved superior across all species and scenarios. Machine learning methods delivered the highest accuracies in certain cases but demonstrated considerable species-specific variability; in contrast, GBLUP consistently provided stable, well-calibrated predictions characterized by low bias.
Furthermore, incremental feature selection of SNPs (Single Nucleotide Polymorphism, the most common type of genetic variation in which a single nucleotide at a specific position in the DNA sequence differs between individuals within a population) enhanced prediction accuracy by 2.8–4.2 percent in three of the species while utilizing only 0.54–9.64 percent of the total available markers. Study results did not report such gain for a low-heritability trait.
Collectively, these findings demonstrate that genomic prediction performance is highly context-specific, highlighting the critical need to simultaneously account for trait genetic architecture, population genetic characteristics, choice of prediction model and strategic marker selection when designing and optimizing genomic selection programs in aquaculture breeding.
Relevance of research findings to the industry
Aquaculture breeding programs increasingly adopt genomic selection to boost genetic gains (often 15–89 percent for growth, higher for resistance) amid disease pressures, feed costs and sustainability goals. This unified benchmark fills a critical gap in fragmented literature, offering practical guidance for model and marker strategy selection.
GBLUP’s reliability and low bias position it as a safe, default choice for routine implementation, especially in lower-heritability or variable environments (e.g., salmon gill disease, bream pasteurellosis). Machine learning’s occasional outperformance suggests value for capturing nonlinearities in high-h² or complex traits (e.g., carp growth, trout survival), though variability requires rigorous validation to prevent poor generalization or overfitting.
SNP feature selection gains (up to 4.2 percent with <10 percent markers) are particularly industry-relevant: reducing genotyping density lowers costs dramatically while preserving or enhancing accuracy – vital for commercial scaling in cost-sensitive sectors or smaller operations. Species with high heritability for specific traits benefit most from advanced models and optimized panels, potentially accelerating profitability, resilience to pathogens and reduced antibiotic reliance.
The context-dependent findings support tailored genomic selection adoption across major species, informing investment in genotyping infrastructure, computational tools, training and reference population expansion. Overall, the results of this study promote evidence-based optimization, enhancing efficiency in global aquaculture breeding.
Aquaculture tourism: An unexpected synergy for the blue economy
Perspectives
This study offers a standardized, cross-species comparison of genomic prediction performance in four key aquaculture species, evaluating GBLUP, various Bayesian methods and machine learning models. Prediction accuracy showed considerable variation across species and traits, driven primarily by trait heritability, population genetic structure and the specific model employed. No single model emerged as the best performer in every situation.
In certain instances, machine learning methods delivered higher accuracy than traditional approaches, though their results exhibited substantial variability depending on the species. In contrast, GBLUP provided consistently stable and well-calibrated predictions with minimal bias. Additionally, applying incremental SNP feature selection enhanced prediction accuracy in multiple species by filtering out low-value or noisy markers, with the degree of improvement tied closely to the underlying genetic architecture of the trait.
Collectively, these findings highlight the substantial differences in genomic prediction effectiveness among species and traits, emphasizing the importance of customizing model choices and marker utilization strategies for different aquaculture breeding contexts.
Overall, authors advocate for broader, more consistent benchmarking efforts to establish reliable guidelines for genomic selection. Looking ahead, promising avenues include assessing additional traits and species (such as shrimp and tilapia), developing multi-trait prediction frameworks and combining genomic data with functional genomics or multi-omics approaches to gain deeper understanding of the underlying biological mechanisms.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-

Darryl Jory, Ph.D.
Editor Emeritus
[103,114,111,46,100,111,111,102,97,101,115,108,97,98,111,108,103,64,121,114,111,106,46,108,121,114,114,97,100]
Tagged With
Related Posts

Health & Welfare
Broodstock management and early hatchery rearing of Arctic charr
Reviewing and summarizing available information on Arctic charr broodstock management, early hatchery rearing and areas of needed research.

Innovation & Investment
AquaGen CEO: Genomics are transforming aquaculture
The CEO of AquaGen knew that the Norwegian research group’s work in genomics was key to the salmon industry’s future. And that was before she even worked there.

Health & Welfare
Mojave yucca extracts are a beneficial phytogenic aquafeed additive
Yucca extracts can enhance farmed fish and shrimp growth and immunity, prevent antioxidative stress and improve culture water quality.

Aquafeeds
Testing alternative diet compositions to offer more sustainable feed formulations for European sea bass
Alternative diets using novel plant and animal-derived protein sources have comparable performance to current commercial sea bass feeds.


![Ad for [membership]](https://www.globalseafood.org/wp-content/uploads/2025/07/membership_web2025_1050x125.gif)