New tool offers accurate, affordable solution to monitoring local biodiversity

A new genomics marker tool has been shown to precisely identify tilapia species and distinguish hybridization between invasive and native tilapia species. In collaboration with the Tanzania Fisheries Research Institute, Roehampton University, Bangor University, the University of Bristol and the University of East Anglia in the United Kingdom, the Earlham Institute led the development of an optimized design based on 96 single nucleotide polymorphisms (SNPs) biomarkers, which proved to be more accurate than microsatellite or morphological identification of interspecific hybrids.
“It’s important to be able to spot these hybrids, yet it’s unreliable to separate them from pure tilapia species by physical characteristics alone,” said lead author Dr. Adam Ciezarek, Postdoctoral Scientist in the Haerty Group at the Earlham Institute. “We have shown that it is possible to identify them using full-genome data. Crucially, we have also demonstrated that a vastly reduced set of 96 SNPs can perform just as well – with greater efficiency and precision, and at a much lower cost.”
The novel SNP markers provide an important resource for assessing broodstock purity in fishery hatcheries, helping to conserve endemic biodiversity. It’s a more affordable and convenient tool that can be used to accurately assess potential farm stocks, as well as survey natural water bodies for evidence of hybridization between tilapia.
In particular, the research team is hopeful that the tool could help develop aquaculture and empower conservation in Tanzania, Africa – a hotspot of natural diversity for tilapia species. At least eight fully endemic Oreochromis species are found in Tanzania and an additional 12 species that are endemic to catchments shared with neighboring countries. Several of these species are adapted to unique environmental conditions (e.g., elevated temperatures, salinity, and pH) and could be of interest for future aquaculture developments.
“Case studies indicate several locations where introduced aquaculture species have become established in the wild, threatening native Oreochromis species in Tanzania,” said lead co-author Prof George Turner, School of Natural Sciences at Bangor University. “At present, native Oreochromis species are poorly characterized, and their conservation could benefit from the identification of purebred populations for protection.”
“Such safeguarding of the wild relatives of farmed species would also protect unique genetic resources that could be used to enhance traits in cultured Oreochromis cichlid strains.”
The peer-reviewed paper, ‘Whole genome resequencing data enables a targeted SNP panel for conservation and aquaculture of Oreochromis cichlid fishes’, was published in Aquaculture. The study was funded by the UKRI Biotechnology and Biological Sciences Council, the Royal Society and the Leverhulme Trust.
Follow the Advocate on Twitter @GSA_Advocate
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Responsible Seafood Advocate
[103,114,111,46,100,111,111,102,97,101,115,108,97,98,111,108,103,64,114,111,116,105,100,101]
Tagged With
Related Posts

Health & Welfare
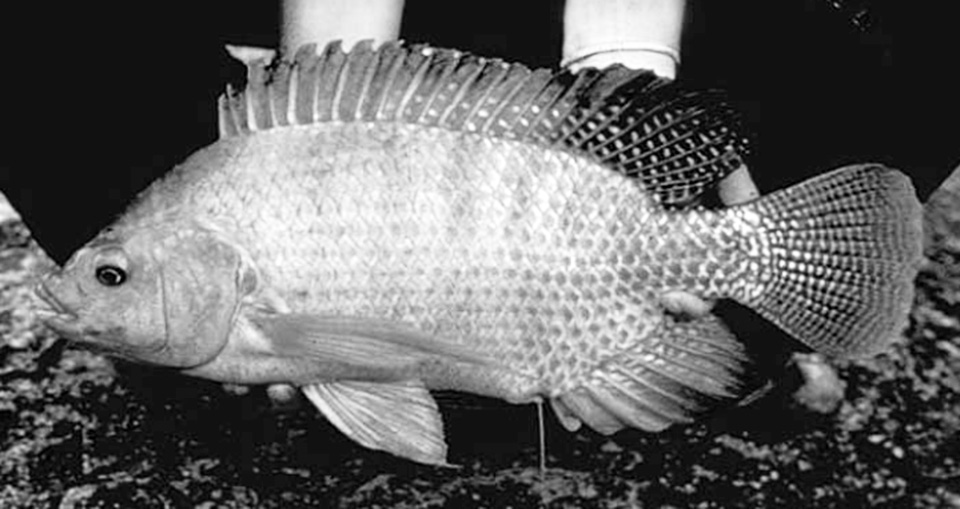
‘Super male’ Nile tilapia outperform hormone-reversed fish in Nicaragua
A recent study compared the growth and survival at different salinity levels of “super male” (YY) tilapia with normal sex O. niloticus reversed to 100 percent phenotypic males.

Health & Welfare
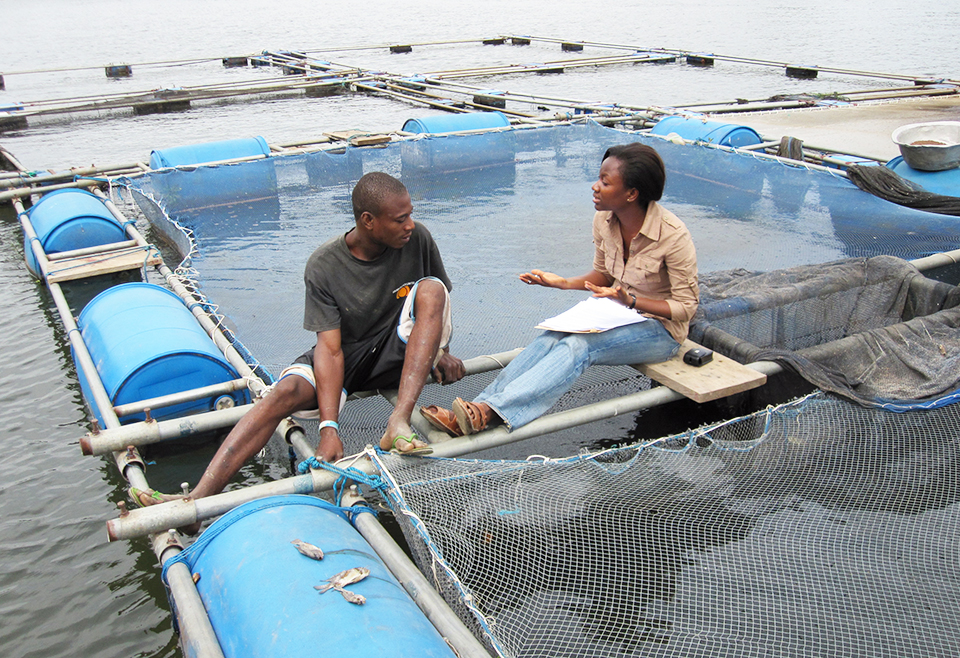
A look at tilapia aquaculture in Ghana
Aquaculture in Ghana has overcome its historic fits and starts and is helping to narrow the gap between domestic seafood production and consumption. Production is based on Nile tilapia.
Health & Welfare
A look at aquaculture genomics
Advances in genomics assist aquaculture science by deepening the understanding of adaptation, physiology and quantitative genetics.

Innovation & Investment
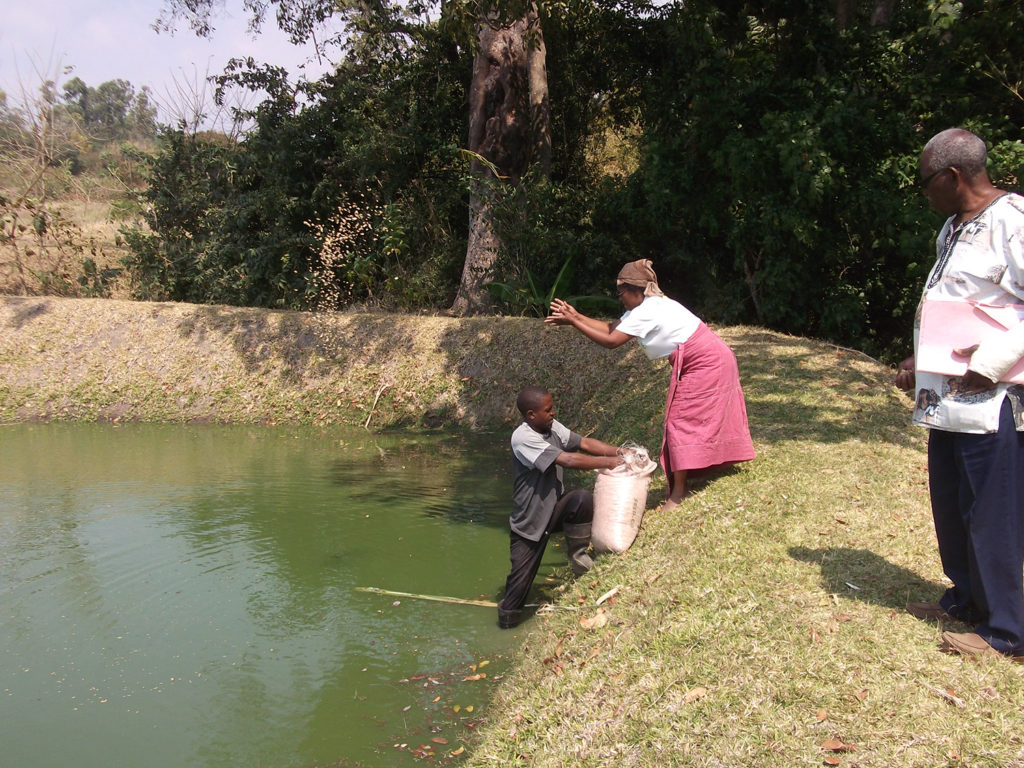
Investing in Africa’s aquaculture future, part 1
What is the future that Africa wants? Views on how to grow aquaculture on the continent vary widely, but no one disputes the notion that food security, food safety, income generation and job creation all stand to benefit.


![Ad for [BSP]](https://www.globalseafood.org/wp-content/uploads/2025/07/BSP_B2B_2025_1050x125.jpg)
