Results show Vibrio population from L. vannamei shrimp’s gut associated with developmental stage and environment
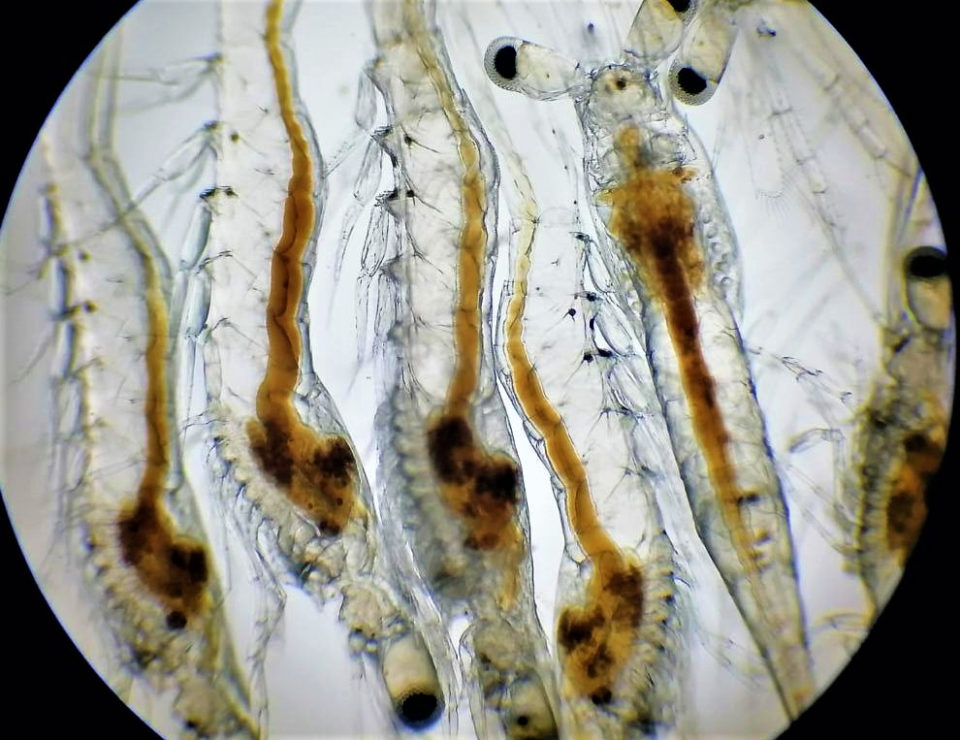
Shrimp diseases caused by opportunistic bacteria are a major problem that can cause considerable losses to the aquaculture industry. The main shrimp diseases are caused by bacterial pathogens mainly from Vibrio species that cause necrosis, slow growth, anorexia and mortality during the shrimp post-larval development.
Research on diseases caused by Vibrio should focus on the biotic interactions between the environment, the host and the pathogenic and non-pathogenic fractions of microbial communities. For instance, Vibrio pathogens can interact with symbiotic communities and the host microbiota [ecological communities of commensal, pathogenic and symbiotic microorganisms found in and on all multicellular organisms] to invade the host’s intestine manipulating its biomechanics to redefine gut communities.
On the whole, understanding the ecological processes implicated in diseases is complicated due to the high diversity of the Vibrio populations, and because the biotic interactions within and between microbial communities modify the disease expression on different levels.
In the same way, the microbial communities could provide guest protection against pathogenic infections because some microorganisms produce antimicrobial complexes. Also, the gut microbiota of shrimp has become more relevant due to our knowledge of its biological interactions. Studying aspects related to the behavior of the gut microbiota may contribute to our understanding of these complex communities.
Despite efforts to improve shrimp production, there are only a few studies based on microbial ecology that link the occurrence of relevant pathogenic microbial groups and the metabolic activities of these groups, causing dysbiosis [a microbial maladaptation or imbalance on or inside the body, like an impaired microbiota] in L. vannamei, which is related to their ontogenetic development.
This article – adapted and summarized from the original publication [Garibay-Valdez, E. et al. 2020. The implication of metabolically active Vibrio spp. in the digestive tract of Litopenaeus vannamei for its post-larval development. Sci Rep 10, 11428] – evaluated the link between the occurrence/abundance of Vibrio populations and bacterial composition in the intestine of Pacific white shrimp (Litopenaeus vannamei) during post-larval ontogenetic development and in its culture water, and the correlation of these with environmental parameters.
https://www.aquaculturealliance.org/advocate/environment-bacterial-community-dynamics-and-white-feces-disease-outbreaks-in-shrimp-ponds/
Study setup
L. vannamei postlarvae (PL5) from the aquaculture farm Parque Acuícola Cruz de Piedra (Guaymas, Sonora, Mexico) were transported in aerated tanks with the same pond water as the farm to the Departamento de Investigaciones Científicas y Tecnológicas de la Universidad de Sonora (DICTUS). A bioassay was conducted with healthy shrimp, each weighing 0.5 ± 0.1 grams, and postlarvae were randomly distributed in the lab recirculation aquaculture system (RAS). Influent seawater was filtered and flowed through a UV lamp for sterilization. The RAS culture units had mechanical aeration and the water salinity was maintained at around 35 ppt with the addition of sterile water to avoid the incorporation of outside bacteria and to compensate for evaporation.
The bioassay started with 40 PL5 randomly stocked into each culture unit, and the experiment lasted 80 days. Throughout the experiment, shrimp were fed twice a day at a rate of 4 percent wet biomass/day using feeding trays with the same formulated feed consisting of commercial grow-out pelleted feed. Samples of intestines were dissected, and water samples were collected for analysis by polymerase chain reaction [a method widely used to rapidly make millions to billions of copies of a specific deoxyribonucleic acid (DNA, molecule carrying genetic instructions for the development, functioning, growth and reproduction of all known organisms) sample, allowing a very small sample of DNA to be amplified it to a large enough amount to study in detail].
The total and metabolically active populations of Vibrio in the digestive tract of shrimp during its postlarval development were analyzed using quantitative PCR (qPCR; to estimate the quantity of starting material in a sample) and reverse transcription qPCR [RT-PCR; measures the amount of a specific RNA (nucleic acid essential in various biological roles in coding, decoding, regulation and expression of genes)] targeting a the 16S rRNA gene sequence [used in reconstructing phylogenies (the evolutionary history and relationships among or within groups of organisms), due to the slow rates of evolution of this region of the gene].
For detailed information on the experimental setup and animal husbandry; collection of intestine and water samples and DNA and RNA extraction; qPCR and RT-qPCR assays; and statistical analyses, refer to the original publication.
https://www.aquaculturealliance.org/advocate/in-vitro-protein-digestibility-of-a-microbial-enhanced-protein-for-juvenile-white-shrimp/
Results and discussion
To evaluate the growth of the L. vannamei in our trial, biometric analyses were performed, demonstrating significant differences based on the developmental stage of the shrimp, with two main stages observed. The first one included animals from days 1 to 40, where their growth was notorious, and the second stage encompassed shrimp that were growing but at a much lower rate, registering no significant differences between days 60 and 80. The first group corresponded to shrimp during the first post-larval developmental stage, while the second group corresponded to shrimp on the sixtieth day of development, when shrimp were considered juveniles.
We estimated the specific growth rate (SGR; the increase in cell mass per unit time) to explain the rate of increase of the L. vannamei population from one sampling day to another, with each interval belonging to a developmental stage. There were significant differences in the SGR between all post-larval stages. From the first stage, the SGR was 5.60 percent per day, followed by the second stage with 4.24 percent per day. At stages III and IV, the SGR of shrimp decreased to 1.36 percent and 0.33 percent per day, respectively, with a global SGR of 2.88 percent per day for the entire bioassay. No significant differences were found between the developmental stages regarding water quality: temperature (24.8 to 25.4 degrees-C), salinity (35.36 to 36.25 percent), DO (6.4 to 6.9 mg per L and pH (8.1 to 8.4).
The shrimp’s nutritional requirements change according to its development and biological behavior. Previous reports demonstrated that the gut enzymatic profile differs between each stage in order to achieve better nutrient absorption. Other researchers showed that the shrimp individual growth rate is associated to its microbiota since organisms of the same age and culture time may have differences in their growth rate, and they present similarities between the gut community assembly according to their growth. In our study, there was a significantly higher growth rate during the postlarval phase compared to the juvenile-adult phase.
Based on qPCR and RT-qPCR analyses, the abundance of the total bacterial community (16S rDNA and 16S rRNA) in the shrimp intestine was demonstrated to be stable from day 20 and onwards. Before that, samples from the control were significantly lower than other culture days. Despite this, some factors including disease, nutrition and environment can regulate the gut microbiota. Our results suggest that a colonization process occurs during the post-larval phase, reaching stability during the juvenile to adult phase, reducing the risk of dysbiosis. However, our results demonstrated that the presence of metabolically active bacteria decreased at culture days 40 and 60, and the amount of this bacteria increased on day 80, with no significant differences concerning days 0 and 20.
The total abundance of the bacterial community in water samples was constant during the bioassay, except for the control (day 0) where the abundance was significantly higher. However, the abundance of metabolically active bacteria was the highest on day 80, and the abundance was significantly higher than days 0, 20 and 60. Previous studies have revealed that both shrimp and its culture water share the same bacterial composition, meaning that shrimp can have similar bacterial assemblages as their culturing media. Also, species richness tends to be higher in water samples than in gut shrimp microbiota.
Previous authors have reported that increases in the Vibrio community or specific bacterial variations in shrimp intestines or water are considered indicators of disease. However, their presence could also be associated with their metabolically active performance, but this is not determined only by their occurrence. Researchers have demonstrated that the proportion of the Vibrio population in shrimp gut microbiota was higher during the nursery stage (2.02 percent) than in the harvest stage (0.64 percent), showing that harvest stage microbiota is more diverse than nursery microbiota.
The results of this study demonstrate that the total relative abundance of Vibrio spp. in shrimp intestines is higher than its metabolic activity. This agrees with other previous studies, where the innate immune system activation of shrimp (Kuruma shrimp, Marsopenaeus japonicus and L. vannamei) was evaluated in response to infections by V. alginolyticus and V. parahaemolyticus, demonstrating that they could activate their defense system in the presence of potentially pathogenic microorganisms, activating genes that have activities related to immune response mechanisms. Therefore, a selective biological pressure exerted by the host on the metabolic activity of these bacteria could be occurring, while stimulating the shrimp’s immune system by functioning as a natural probiotic necessary for the life cycle of the host.
Perspectives
Our results provide an overview of the interactions between the intestinal microbiota of L. vannamei shrimp and its environment, emphasizing the ratio and correlation between the total and metabolically active Vibrio. Both the total and active populations of Vibrio indicated that there is colonization of this bacterium in the white shrimp intestine throughout its post-larval development, but with low metabolic activity. Therefore, Vibrio is part of the intestinal microbiota of the shrimp, and it could be playing dualistic role, involved in a symbiotic relationship between the organism and the bacterial community structure of the intestinal microbiota, while acting as an opportunistic pathogen under certain circumstances.
Our data suggest that the rest of the microbiota could probably regulate Vibrio activity, possibly establishing a coexistence mechanism that does not affect the host, providing conditions for the microbial community to thrive. In contrast, the abundance of the metabolically active Vibrio population was greater in crop water, despite its low total abundance. Finally, our results demonstrate that Vibrio can be a common and inclusive a dominant member of the white shrimp microbiota, but its activity could be regulated by the rest of the microbiota and by the shrimp.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Estefanía Garibay-Valdez, Ph.D.
Centro de Investigación en Alimentos y Desarrollo A.C (CIAD)
Carretera a La Victoria S/N, CP. 83304, Hermosillo, Sonora, Mexico -
Luis Rafael Martínez-Córdova, Ph.D.
Departamento de Investigaciones Científicas y Tecnológicas (DICTUS)
Universidad de Sonora, Blvd. Luis Donaldo Colosio S/N, CP. 83000, Hermosillo, Sonora, Mexico -
Marco A. López-Torres
Departamento de Investigaciones Científicas y Tecnológicas (DICTUS)
Universidad de Sonora, Blvd. Luis Donaldo Colosio S/N, CP. 83000, Hermosillo, Sonora, Mexico -
F. Javier Almendariz-Tapia, Ph.D.
Departamento de Ingeniería Química y Metalurgia
Universidad de Sonora, Blvd. Luis Donaldo Colosio S/N, CP. 83000, Hermosillo, Sonora, Mexico -
Marcel Martínez-Porchas, Ph.D.
Centro de Investigación en Alimentos y Desarrollo A.C (CIAD)
Carretera a La Victoria S/N, CP. 83304, Hermosillo, Sonora, Mexico -
Kadiya Calderón, Ph.D.
Corresponding author
Departamento de Investigaciones Científicas y Tecnológicas (DICTUS)
Universidad de Sonora, Blvd. Luis Donaldo Colosio S/N, CP. 83000, Hermosillo, Sonora, Mexico
Tagged With
Related Posts

Health & Welfare
Assessing effects of long-term pH stress to Pacific white shrimp juveniles
Evaluating the effects of chronic pH stress on juvenile Pacific white shrimp, study results suggest adverse affects on gut microbiota functions.

Health & Welfare
Intestinal bacteriome of Pacific white shrimp in biofloc, clear water systems
Global overview of the intestinal bacteriome of Pacific white shrimp in biofloc and clear water systems, and after an oral challenge by white spot syndrome virus (WSSV).
Health & Welfare
Impact on gut microbiota health in L. vannamei postlarvae after salinity reduction treatment
Study evaluates effects on Pacific white shrimp gut microbiota after salinity reduction, addition of Bdellovibrio and like organisms (BALOs).

Health & Welfare
Probiotics benefit Pacific white shrimp challenged with AHPND
A study was conducted to measure the effects of commercial probiotics on Pacific white shrimp in a standardized AHPND challenge model under controlled laboratory conditions. Results show that the probiotics treatments by themselves have beneficial effects, such as higher survival and histological signs of hepatopancreas regeneration.


