Aquacultured and released shrimp comprised 2.67% of the total recaptures and did not produce significant changes in the genetic structure of wild populations
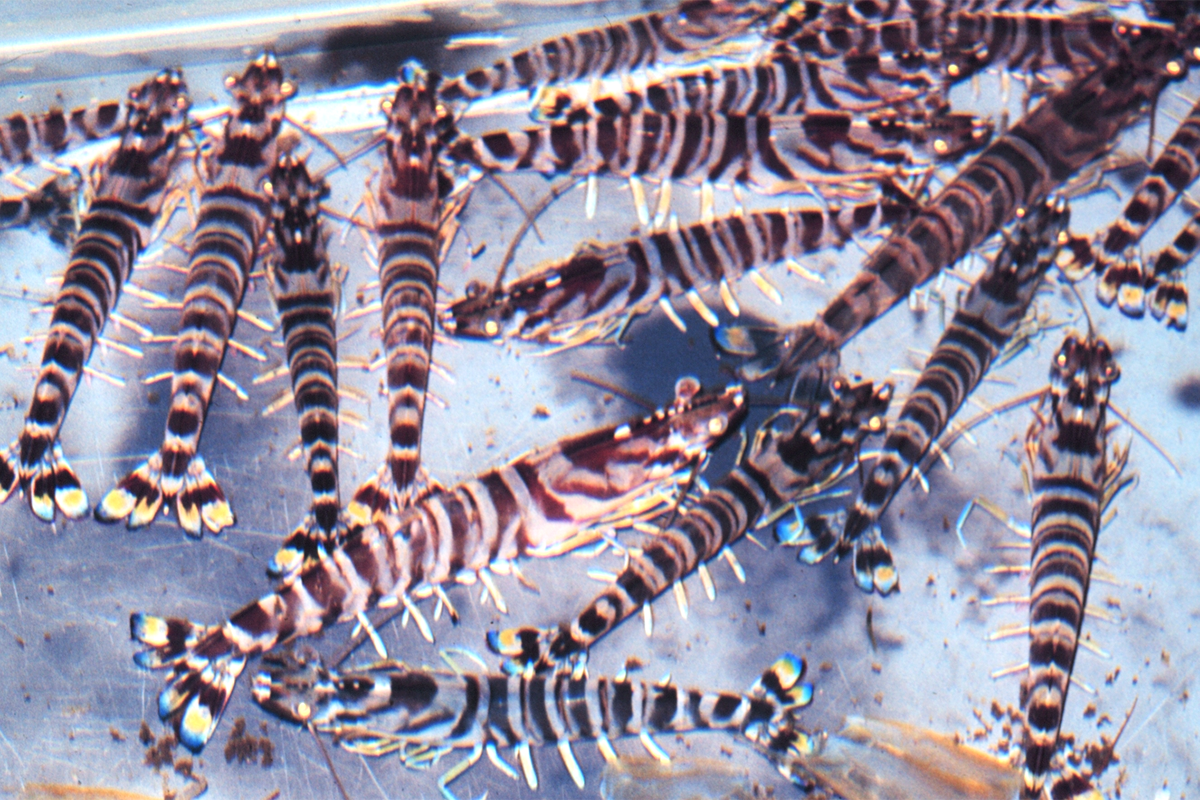

The Kuruma shrimp (Penaeus japonicus) is an important commercial species in the Indo-West Pacific, supporting valuable fisheries and also commercial aquaculture. To restore declining natural populations of Kuruma shrimp in the Shandong Peninsula region of China, stock enhancement programs were established and benefits were shown by increased fisheries yields. However, despite the scale of the stock enhancement increase, there are still few studies on genetic monitoring and genetic diversity before and after stock enhancement activities.
Stock enhancement is an effective way to facilitate the recovery of fishery resources, where aquacultured young animals are released into natural waters for the purpose of restoring or increasing the number in natural stocks to achieve the rapid replenishment of these fishery resources.
Studies indicate that stock enhancement is one of the more effective measures to support wild populations of P. japonicus. But when genetic exchange occurs between the released population and the wild population, it may cause changes in the genetic structure of the population within the natural environment; therefore, it is particularly important to study the genetic diversity of the P. japonicus population before and after stock enhancement.
Microsatellites (microsatellite DNA) are composed of repeated DNA base pairs and have many advantages as molecular markers compared with other genetic markers. They can be used for population genetics, pedigree analysis and population genetic-map construction. Microsatellite markers are also important in the aquaculture sector, contributing to the conservation and improvement of cultured species through their genetic analysis, helping to understand species diversity, phylogeny, evolution and development, and providing a basis for the sustainable conservation and development of aquaculture.
This article – summarized from the original publication (Zhang, M. et al. 2023. Microsatellite-Marker-Based Evaluation of Stock Enhancement for Kuruma Prawn Penaeus japonicus in Beibu Gulf, South China Sea. Fishes 2023, 8(12), 568) – reports the results of research to evaluate the effect of stock enhancement on P. japonicus in the Gulf of Beibu, South China Sea, and to investigate whether stock enhancement has genetically affected the natural population of this shrimp species in this area.
Study setup
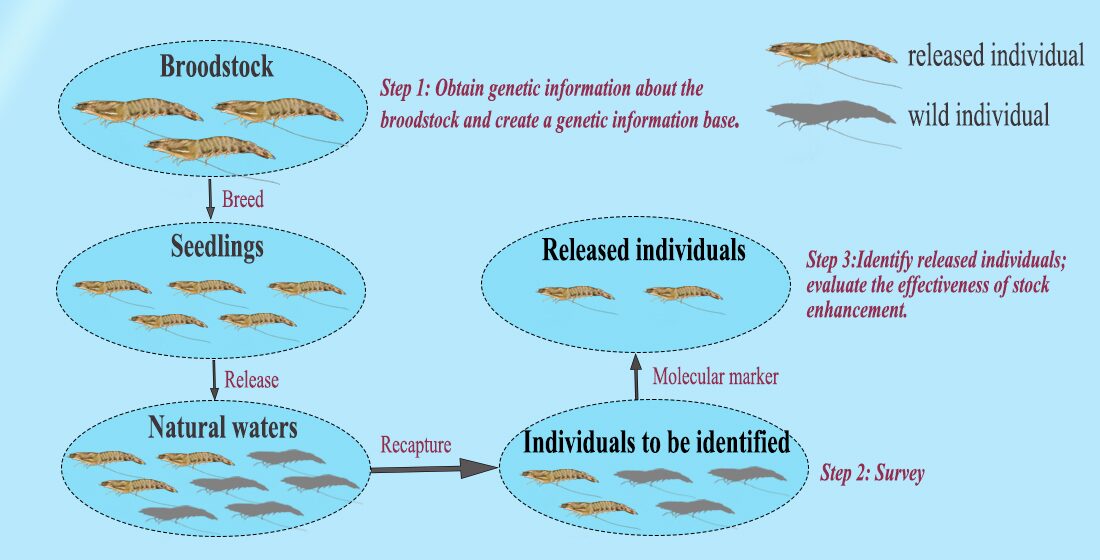
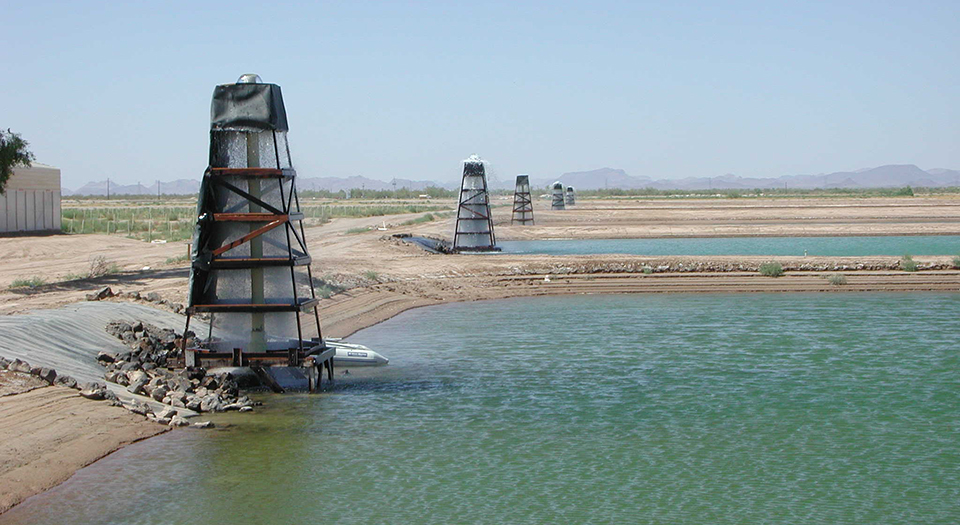
Wild, mature P. japonicus females were collected for artificial breeding in the waters near Beihai, South China Sea, and approximately 5 million offspring shrimp were successfully produced. After morphological data and tail tissue samples were collected and preserved, these animals were released in the stock enhancement area. After release, monthly recaptures of P. japonicus were conducted for four months at five sampling sites. Morphological measurements were taken on the recaptured shrimp and tail muscle samples collected for analyses and comparisons.
We used microsatellite molecular markers to identify released individuals among recaptured animals and calculated the recapture rate of P. japonicus to preliminarily assess the release of these aquacultured shrimp in the Bay of Beibu, China. The genetic differences between the broodstock, released and recaptured populations of P. japonicus were also determined to analyze the genetic effects of stock enhancement.
For detailed information on the study design, animal rearing and release, data collection and analyses, refer to the original publication.
Results and discussion
To help restore the resources of P. japonicus in the waters of the South China Sea and the Gulf of Beibu, aquacultured young shrimp were released in this study of resource restoration and the effect of stock enhancement was assessed by using microsatellite molecular markers.
The assessment of stock enhancement effects is an important measure for the effective management of stock enhancement, and the assessment results can be used as a basis for improving the technical methods of stock enhancement and to provide better technical support for the recovery of fishery resources. The use of molecular marker technology has become one of the most effective means for the accurate and long-term assessment of stock enhancement effects.
Based on the data on microsatellite loci (tracts of repetitive DNA) assessed, we conclude that the results of this study are consistent with those of previous reports and indicate that the P. japonicus populations in this study had a high level of genetic diversity.
In this study, we evaluated the stock enhancement activity of P. japonicus and studied the shrimp of this species recaptured during four months after release based on microsatellite markers to assess the stock enhancement effect. Among 487 recaptured individuals, 13 individuals were identified as released individuals, resulting in a recapture rate of 2.67 percent. In a similar, previous study in Japan, the recapture rates of P. japonicus used for stock enhancement were reported to range from approximately 0.0 percent to 22.1 percent in 40 cases but exceeded 10 percent in only two cases. This result shows that the recapture rate after stock enhancement in Japan was mostly below 10 percent, consistent with the results of our study.
Stock enhancement may have an impact on the genetic characteristics of the target marine species, and the aquacultured animals released into the natural environment may alter the genetic diversity of the wild population; e.g., intraspecific competition occurs between the released population and the natural population, and if the released population dominates in events of intraspecific competition, the size of the natural population will decrease and a corresponding reduction in genetic diversity will occur.
We studied the genetic diversity among the broodstock population, the released population and the recaptured population. The results for the three P. japonicus populations in our study for the parameters analyzed indicated that these three P. japonicus populations had a high level of genetic diversity.
The levels of genetic differentiation among populations are usually measured by the genetic distance and genetic differentiation coefficient. Our data indicated that there was significant genetic differentiation among the broodstock population, the released population and the recaptured population. If the genetic differentiation between released and natural populations is significant, it may lead to changes in the genetic structure of the P. japonicus populations in natural waters, which should be avoided. This may be because a sufficient number of parents from the target waters were not used to breed the shrimp seedlings, which diminished the similarity with the genetic background of the wild populations. More wild populations from the target waters should be introduced as broodstock to increase the diversity of released populations and to prevent greater genetic differentiation from natural populations.
A final consideration is the potential introduction of novel pathogens into the natural environment via the released breeding population, which may result in higher mortality in the wild population due to a lack of immunity to any novel pathogens, which will lead to a reduction in the size of the wild population and, consequently, a reduction in the level of genetic diversity in the natural population.
Altogether, our results showed that the released P. japonicus population did not have a significant demographic or genetic impact, evaluated with microsatellite markers, on the natural population. However, if the resources and ecological balance of P. japonicus are to be maintained in the long term, the continuous genetic monitoring of natural populations using more molecular markers is required. More attention should be given to the possible effects on the genetic aspects of the natural population to achieve the sustainable use of its resources.
Perspectives
In this study, we used microsatellite molecular markers to identify released individuals among recaptured specimens and calculated the recapture rate of P. japonicus to preliminarily assess the release of these shrimp in the Bay of Beibu, China. The genetic differences between the broodstock, the released population and the recaptured population of P. japonicus were also determined to analyze the genetic effects of stock enhancement. Among the 487 recaptured individuals, 13 individuals were identified as released individuals, resulting in a recapture rate of 2.67 percent.
The results of this study together showed that the released P. japonicus population did not have a significant genetic impact, evaluated with microsatellite markers, on the natural population. However, if the resources and ecological balance of P. japonicus are to be maintained in the long term, we recommend the continuous genetic monitoring of natural populations using more molecular markers.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Dr. Dianrong Sun
Corresponding author
South China Sea Fisheries Research Institute, Chinese Academy of Fisheries Sciences; and Key Laboratory of Marine Ranching, Ministry of Agriculture Rural Affairs, Guangzhou 510300, China[110,99,46,99,97,46,105,114,102,115,99,115,64,51,55,110,117,115,114,100]
Tagged With
Related Posts

Intelligence
Farming kuruma shrimp in Japan
Kuruma shrimp (Penaeus japonicus) are produced on a small scale. This species can tolerate transportation over long distances and without water.

Health & Welfare
How RNAi-based therapies combat viral shrimp diseases
By precisely targeting viral pathogens, RNAi has the potential to provide an environmentally friendly solution to combatting viral shrimp diseases.

Health & Welfare
Phospholipids: Essential nutrients for shrimp
Phospholipids play important roles in maintaining normal cell structure and functions and are sources of second messengers in cell signaling.

Innovation & Investment
A novel chromosome-level genome assembly of the Pacific white shrimp
A genome assembly of L. vannamei provides valuable resources for future genetic research, breeding and improvement of traits for aquaculture.


![Ad for [Aquademia]](https://www.globalseafood.org/wp-content/uploads/2025/07/aquademia_web2025_1050x125.gif)
