Method has advantages for diagnosis over common histological techniques
Aquaculture of blue crabs (Callinectes sapidus) is being developed in the Chesapeake Bay between Virginia and Maryland, United States, to reduce the impact of overfishing. The growing demand for both hard- and soft-shell crabs has led to overfishing, with an estimated 75 percent of the bay’s adult blue crab stock removed each year. This has resulted in dramatic reductions of blue crab populations.
Historically, blue crab aquaculture in Chesapeake Bay has been relatively primitive and based on the capture and fattening of juvenile crabs collected from the wild. Current efforts are focused on the development of hatchery technology that will increase the availability of juvenile blue crabs for both aquaculture and stock enhancement.
Recent success in the hatchery technology has shown that blue crabs can be mass produced year round, which increases the prospects for commercial production. Attention to blue crab diseases, especially those caused by marine viruses, will undoubtedly become increasingly important as the industry develops.
Blue crab reovirus
A Callinectes sapidus reovirus (CsRV) found in blue crabs captured from Chesapeake Bay in 2005 was present in over 50 percent of dead/moribund soft-shell crabs and 5 percent of the wild crab populations sampled. As noted in 2010 by Holly Bowers and co-authors in Diseases of Aquatic Organisms, this virus replicates in the cytoplasm of infected cells. It has an icosahedral shape with a 55-nm diameter and contains a genome consisting of 12 segments of dsRNA.
Histopathological characterization
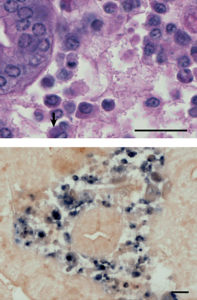
Histological examination of the affected crabs showed the presence of eosinophilic to basophilic cytoplasmic inclusions in both hepatopancreas and gill tissues. In the hepatopancreas, inclusions were found in the cells of connective tissue and hemocytes in hemal sinus. These spherical to pleomorphic inclusions were present both singularly and in clusters. In the advanced of infection, the cytoplasmic area showed an increased volume, and the inclusions became less intense in color with a granular appearance.
In the gill filaments, the inclusions were also found in hemocytes. In addition, focal inflammatory lesions were present, which were formed by repeated layering of fibrocytic cells. The core of these lesions consisted of infected hemocytes which engulfed necrotic cell debris.
Total RNA was extracted from hemolymph samples drawn from the moribund crabs, and the cDNA was synthesized for construction of a cDNA library. One clone (CsRV-28) was found to have a significant match with a reovirus that infects mud crabs (Scylla serrata). The matched region is part of the guanylytransferase gene, and there was a 95 percent similarity in amino acid sequence.
In situ detection
The CsRV-28 clone was used for generation of a digoxigenin-labeled probe for in situ hybridization. This probe reacted to cytoplasmic inclusions in the hemocytes of hepatopancreas and gill tissues. No reaction was seen in any of the tissues prepared from uninfected crabs. Thus, the in situ hybridization was effective as a diagnostic procedure.
This method has several advantages for diagnosis over commonly used histological techniques. It provides a high specificity in that the positive reaction occurs only with the target virus and thus can distinguish among viruses that are morphologically similar. In addition, the method is relatively easy to perform, rapid and suitable for routine use by diagnostic laboratories.
(Editor’s Note: This article was originally published in the November/December 2011 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Authors
-
Kathy F.J. Tang, Ph.D.
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA[117,100,101,46,97,110,111,122,105,114,97,46,117,64,117,121,106,103,110,101,102]
-
Carlos R. Pantoja, Ph.D.
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA -
Rita M. Redman
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA -

Donald V. Lightner, Ph.D.
Department of Veterinary Science and Microbiology
University of Arizona
Tucson, Arizona 85721 USA
Tagged With
Related Posts

Responsibility
Examining copper use in aquaculture
Copper is used for control of the blue-green algae responsible for off-flavors in aquaculture animals, treating diseases and parasites, and avoiding cage net fouling. Although copper is an essential nutrient for plants and animals, an excess can negatively affect the environment and human health.

Health & Welfare
Flatfish conditioning for stock enhancement
Acclimation cages, the most implemented conditioning strategy for flatfish, are inexpensive and effective, but require site-specific adjustment. Live or lifelike diets show great potential in rearing stocked flatfish.

Health & Welfare
Malaysia shrimp project scales up for production in biosecure biofloc modules
A large-scale integrated shrimp aquaculture park (iSHARP) project in Malaysia is approaching completion of its first phase. Biosecurity is a priority at iSHARP. The design of each unit allows individual modules or ponds to be “locked down” to prevent disease from spreading.

Health & Welfare
Recent developments in biofloc technology
Combining biofloc technology with bio-secure modular shrimp culture can make operations more sustainable and economically viable. For optimized biofloc production, lined ponds and reservoirs and high stocking densities are essential. Paddlewheel aerators keep dissolved-oxygen levels high.


![Ad for [Aquademia]](https://www.globalseafood.org/wp-content/uploads/2025/07/aquademia_web2025_1050x125.gif)
