Linkage maps constructed for major aquaculture fish and shellfish species

The human genome project marked the start of the so-called genomics revolution. In the last 20 some years, genomics has undoubtedly changed the way scientific research is conducted in the life science areas.
In the aquaculture sector, major genome research was initiated in 1997, when a group of scientists met in Dartmouth, Mass., USA, to discuss aquaculture genome initiatives. In the years after, the aquaculture genome project was initially classified as a regional multistate project. In 2003, the aquaculture group joined the National Animal Genome Project of the United States Department of Agriculture as a part of national project NRSP-8.
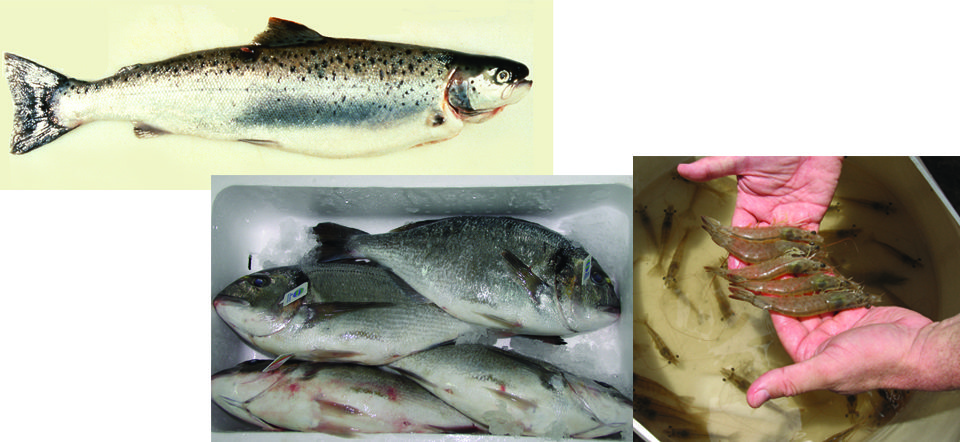
The objectives for the first five years of the national project were to develop DNA markers, construct genetic and physical maps of aquaculture species and align the maps for studies of performance and production traits. Six major species groups were included: catfish, Atlantic salmon and rainbow trout, tilapia, shrimp, oysters and striped bass.
Major progress in aquaculture genomics
Major progress has been made in aquaculture genomics in spite of the late start. To date, framework genetic linkage maps with several hundred DNA markers have been constructed for major aquaculture fish and shellfish species.
Genetic linkage maps are developed based on inheritance and segregation of DNA markers. Markers on different chromosomes segregate independently. Markers on the same chromosome tend to be inherited as a unit, which marks their linkage. However, markers far apart on the same chromosome tend to segregate because of recombination. The recombination frequency depends on the distances between markers, which define the genetic distances on the linkage map.
Genetic maps are typically useful for studies of performance traits. For instance, DNA markers tightly linked with disease resistance genes cosegregate with resistance phenotypes. These markers can therefore be used to mark the presence of disease resistance genes. In spite of being very useful for studies of performance traits, linkage maps do not provide information or material basis for DNA sequences or clones containing the genes related to performance traits. Such information is included in physical maps of the genome.
Physical maps are most often developed in the form of bacterial artificial chromosome (BAC) libraries. BAC libraries contain multiple copies of the genome that is first randomly broken into large segments. Segments of overlapping regions from multiple copies of the genome can be organized in a linear fashion to cover the entire genome. The overlapping arrays of genomic DNA are called the BAC-based physical map.
Once a gene is mapped in a genomic region, the corresponding DNA segments can be isolated from the library for detailed analysis. To date, BAC-based physical maps have been constructed for Atlantic salmon, tilapia and channel catfish, and those of rainbow trout and oysters are under construction.
The genetic maps and physical maps are most useful when they are integrated and aligned. Integration of genetic linkage and physical maps are under way for Atlantic salmon and catfish.
Gene discovery, identification
Gene discovery efforts have been made in various aquaculture species. Typically, a fish can have 20,000 to 30,000 genes distributed in the genome. Despite this large number, the genes represent typically less than 5 percent of the total genome. To identify most of the genes from a species can therefore be quite difficult.
One of the most efficient approaches for gene discovery and identification is to directly sequence the complementary DNA (cDNA) libraries, generating short sequences of expressed genes. These short sequences are called expressed sequence tags (ESTs). Large numbers of ESTs have been generated for catfish, Atlantic salmon, rainbow trout, cod, tilapia, oysters and shrimp (Table 1). Detailed sequence analysis and assembly of ESTs would allow the number of genes represented by the ESTs to be revealed. For instance, the catfish ESTs currently available represent approximately 29,000 unique gene sequences.
Liu, Expressed sequence tags, Table 1
| Species | ESTs Available |
|---|---|
| Zebrafish (Danio rerio) | 1,379,829 |
| Atlantic salmon (Salmo salar) | 433,337 |
| Channel catfish (Ictalurus punctatus) | 300,678 |
| Three-spined stickleback (Gasterosteus aculeatus) | 276,992 |
| Rainbow trout (Oncorhynchus mykiss) | 260,887 |
| Atlantic cod (Gadus morhua) | 183,047 |
| White shrimp (Litopenaeus vannamei) | 155,411 |
| Blue catfish (Ictalurus furcatus) | 139,475 |
| Sea lamprey (Petromyzon marinus) | 120,731 |
| Seabream (Sparus aurata) | 50,930 |
| Seabass (Dicentrarchus labrax) | 37,962 |
| Common carp (Cyprinus carpio) | 32,046 |
| Pacific oyster (Crassostrea gigas) | 29,018 |
| Eastern oyster (Crassostrea virginica) | 14,560 |
| Chinese shrimp (Fenneropenaeus chinensis) | 10,446 |
| Japanese flounder (Paralichthys olivaceus) | 8,822 |
| Tiger shrimp (Penaeus monodon) | 8,398 |
| Nile tilapia (Oreochromis niloticus) | 294 |
Microarrays
ESTs are the material basis for cDNA-based microarray technology. Microarrays are sometimes referred to as gene chips. As the name reflects, they are small, but contain a large number of genes on a microscopic glass slide. Typically, over 20,000 genes can be printed on each slide.
Microarrays are useful for gene expression analysis, especially for the expression of tens of thousands of genes before and after any sort of treatment. For instance, disease infection may trigger the expression change of hundreds of genes, and the use of microarray technology would allow the differentially expressed genes to be readily detected. Microarray technology has been developed for the analysis of genome expression in salmonids, catfish, oysters and shrimp.
The genome landscape is characterized by repeat structures. The structures are well studied in catfish, salmonids and several other aquaculture species, laying the grounds for whole genome sequencing. A draft genome sequence is being produced for tilapia, and most recently, whole genome sequencing is being conducted for Atlantic salmon. White papers have been produced that advocate whole genome sequencing of rainbow trout, catfish, oysters and shrimp.
Challenges, advantages
In spite of the above progress, aquaculture genomics is facing great challenges. The greatest challenge is not technical, but financial. As over 300 species are involved in aquaculture, genome studies of aquaculture species require great efficiency.
Genome studies of aquaculture species, however, also have many advantages. The high fecundity of many species allows production of large full-sib and half-sib families suitable for analysis of performance traits. Large resource families allow heavy selection pressure to be applied, which not only enhances the likelihood of genetic linkage detection, but also allows rapid progress in breeding programs.
As the focus of aquaculture genomics should be on performance trait-related issues, efficient and accurate phenotypic evaluations and genotyping systems are crucially important. To meet these challenges, greater efforts in phenomics must include the development of biochemical and molecular indicators of performance traits. Efficient marker platforms must be developed, especially the single nucleotide polymorphism (SNP) marker platforms.
Research must continue the course of generating draft genome sequences for major agricultural and aquaculture species, yet much can be accomplished using existing genome resources and technologies such as 454 and Solexa sequencing, which are often referred to as next-generation technologies.
The efficient SNP platforms, when coupled to the use of large resource families, should yield rapid progress in aquaculture genomics for practical applications and genetic enhancement using marker-assisted selection and other related technologies. Genome research is fundamental basic research, but its application to aquaculture will provide unprecedented accuracy for genetic selection of performance and production traits.
(Editor’s Note: This article was originally published in the November/December 2008 print edition of the Global Aquaculture Advocate.)
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Zhanjiang Liu, Ph.D.
The Fish Molecular Genetics and Biotechnology Laboratory
Department of Fisheries and Allied Aquacultures
Program of Cell and Molecular Biosciences
Aquatic Genomics Unit
Auburn University
Auburn, Alabama 36849 USA[117,100,101,46,110,114,117,98,117,97,64,110,97,104,122,117,105,108]
Tagged With
Related Posts

Innovation & Investment
AquaGen CEO: Genomics are transforming aquaculture
The CEO of AquaGen knew that the Norwegian research group’s work in genomics was key to the salmon industry’s future. And that was before she even worked there.
Health & Welfare
Animal health giants have sea lice in their crosshairs
Alltech and Benchmark have been working on the next generation of sea lice solutions and believe they have new products that can help salmon farmers win.

Health & Welfare
Big shoes to fill: Dhar takes reins at shrimp pathology laboratory
Arun Dhar, Ph.D. will attempt to fill the “big shoes” of Dr. Donald Lightner at the University of Arizona’s Aquaculture Pathology Laboratory, where the shrimp disease EMS was diagnosed.

Aquafeeds
Canada eagerly looks to camelina oil as a fish oil alternative
Grown worldwide, Camelina sativa is a tiny seed used for human consumption that also has big potential for driving the “green” economy.


![Ad for [membership]](https://www.globalseafood.org/wp-content/uploads/2025/07/membership_web2025_1050x125.gif)
