Polymerase chain reaction is a very powerful molecular tool, but what does a positive sample actually mean?
Polymerase chain reaction (PCR) is a very powerful molecular tool that allows the detection of very low levels of deoxyribonucleic acid, or DNA (the genetic code or hereditary material of organisms). At the core of this method is an enzyme that amplifies the amount of DNA present. PCR is highly sensitive and specific, and once a presumptive organism (typically bacteria or viruses, although it can detect DNA of any organism) of interest has been identified, the DNA is sequenced and specific sequences that are likely unique to the organism are characterized.
These nucleotide (the building blocks of DNA) sequences form the basis of a primer. The primer is a template for the reaction. A DNA sample from the organism of interest is mixed with the primer. The primer binds to the homologous (each of the four bases, the building blocks of DNA that pair specifically with each other) DNA sequence if it is present. PCR can be used to determine if a pathogen is present, i.e., a simple yes or no, with no indications of how much is present, or it can be used in a quantitative manner.
PCR technology has revolutionized our understanding of the world around us. We can find potential pathogens at very low levels as well as determine that an infectious process is taking place by quantifying how the levels of the organism of interest change with time. Unfortunately, pseudoscience permeates all facets of most activities that humans engage in, including aquaculture, and PCR is widely misused and poorly understood. This can cause serious problems in terms of false positives, false negatives or even mixed reactions that mislead the user of the technology.
As with most chemical reactions, the conditions under which the PCR tests take place are exacting and even minor variations can impact the outcome. Improper controls, contamination of samples and of reagents, failure to abide by thermal cycling requirements, etc. can all result in test results of questionable value. This is one reason why labs that conduct these tests routinely should be audited by third parties to verify that they are indeed measuring accurately and consistently, as it is too easy to be wrong.
Despite efforts to educate the end users about the real value of PCR, this has not been universally successful. Most fail to understand that this is a tool and not a solution. As the tool has evolved, it has found a significant niche in aquaculture and fisheries. Here I will discuss some of the common widespread fallacies that directly impact the potential value of the PCR technology.
https://www.aquaculturealliance.org/advocate/effect-of-a-commercial-parabiotic-on-shrimp-production/
Fallacy No. 1: Positive results are always real
Properly constructed primers are specific for a given organism. While it is expected that the primer sequence is unique, if it is not unique the primer can cross-react with the DNA from other organisms. Every effort is made to minimize this by focusing on genes that are thought to be unique. For some pathogens, such as the etiologic agent of Infectious Hypodermal and Haematopoietic Necrosis Virus (IHHNV) it is not straightforward. Shrimp are known to incorporate pieces of viral DNA into their genome (all the genetic material).
The primer sequence must be directed against DNA sequences that are consistent with infection and not with incorporation into the hosts’ genomes. The nature of the PCR technology is such that it is possible to engineer primer sequences that are not found in any other organisms. That said, the specter of cross-reactivity resulting in misleading data is all too real. Although appropriate controls are run to ensure that this is not the case, it does not mean that it cannot happen. In the case of IHHNV, if we are looking for carriers that contain virus that can be spread, we have to make sure that the primer sequences are consistent with finding the virus and not the non-infective inclusion.
Fallacy No. 2: Screening of populations using PCR is a statistical tool that is well suited for determining the absence of primer reactive DNA
This technology is widely used for testing large populations. Unfortunately, when used by itself it can lead to false negatives. It is best suited for testing individuals to ascertain carrier status and the potential for specific organisms to be present that could lead to acute disease.
A good example of this is testing for the presence of the virus that causes White Spot Syndrome Virus (WSSV). A major route of transmission of this disease is through carriers. Shrimp broodstock carry the virus and pass it to shrimp postlarvae (PL), which in turn carry it into the production environment. For most highly virulent pathogens there is ZERO tolerance. If they are present and the conditions are consistent with allowing the virus to proliferate, all it takes is a very small number (one) of carriers.
The American Fisheries Society publishes its blue book on fish health to help aquaculturists deal with the myriad disease issues that may affect them. It lays out the statistical basis for sampling populations. Unfortunately, these are not suitable for use in shrimp where high rates of fecundity can result in half a million or more eggs from a single spawn.
The sampling must be done at random, and the technology must be 100 percent accurate. But in large populations, the greatest degree of accuracy is 98 percent, which means that 2 percent of the population can still be carrying a pathogen. Out of a million shrimp PLs, 20,000 could still be carrying the pathogen and PCR testing could tell you that there was nothing. Of course, many times the sampling is not random and the test itself is prone to errors when not conducted properly.
Many companies test broodstock and PLs routinely for a number of pathogens, typically those that the World Organisation for Animal Health (OIE) dictates. Because of the cost of running PCR tests and the lack of regulatory oversight, only a small subsample of animals is typically tested. The end result is that pathogens routinely enter susceptible populations. The use of PCR cannot and does not protect the animals.
Fallacy No. 3: PCR is suitable for testing for the presence of any presumptive pathogen
Not all pathogens are present in all tissues. If the level of infection is very low, then it is important to sample those tissues that are known to be targeted by the pathogen. Furthermore, for some pathogens merely taking a sample is not good enough. Using WSSV as an example once more, this virus replicates very poorly at water temperatures above about 31 degrees-C. For the most part, unless there are serious stressors in play and overt evidence of infection, the viral loads in animals held in warmer water are too low to be detected via PCR.
Even when one focuses on target tissues, the virus can be dormant and finding it is like looking for a needle in a haystack. Animals to be tested need to be held under conditions that will force viral expression. Gradually lowering the water temperature to the mid 20 degrees-C and holding the animals for a few days is the only way to be sure that the virus is not present in a population when testing using PCR. If this is not done, then one cannot state that the population of interest is free of WSSV. In fact, there are many instances where animals are being claimed to be free of WSSV when the virus is dormant, and which have resulted in massive disease outbreaks.
Fallacy No. 4: You can use PCR to establish that a population is SPF
The concept of specific pathogen free (SPF) is important. If one takes the definition literally, it means that a given pathogen is not present in the stocks tested. However, if all one is doing is blue book levels of testing, the population should not be considered to be SPF without some additional background.
PCR by itself cannot allow one to declare a population of animals as being SPF. SPF is a process and not the result of limited testing of a group of animals. The only way one can be sure that a population is SPF, short of testing each animal individually (broodstock) under the appropriate conditions is via their history. This is not a widespread practice and many farmers find themselves in the midst of massive die offs that are a direct result of PLs carrying the virus.
The use of properly constructed nucleus breeding centers (NBCs) along with PCR screening and animal histories is realistically the best way to ensure that a population is indeed SPF. The animals need to be held in a manner that prevents contamination from live feeds, from inadequately screened animals, from poorly designed holding facilities and other considerations. NBCs should never be open – animals can leave the facility, but they cannot re-enter.
Perspectives
PCR testing is a valuable tool when used correctly. A positive result is important while a negative result is less so. A negative result on a subsample of a large population in conjunction with no follow-up and the lack of use of an appropriately engineered NBC can lead to a disaster, and unfortunately it has and will continue to do so. This does not mean that PCR has no value, but that relying on it solely is not consistent with adequate levels of biosecurity and with true sustainability.
Now that you've reached the end of the article ...
… please consider supporting GSA’s mission to advance responsible seafood practices through education, advocacy and third-party assurances. The Advocate aims to document the evolution of responsible seafood practices and share the expansive knowledge of our vast network of contributors.
By becoming a Global Seafood Alliance member, you’re ensuring that all of the pre-competitive work we do through member benefits, resources and events can continue. Individual membership costs just $50 a year.
Not a GSA member? Join us.
Author
-
Stephen G. Newman, Ph.D.
President and CEO
Aquaintech Inc. – “Biotechnology Benefiting Aquaculture”
Lynnwood, WA 98037 USA[109,111,99,46,104,99,101,116,45,110,105,45,97,117,113,97,64,109,119,101,110,103,115]
Tagged With
Related Posts

Health & Welfare
The unmet promise of pondside PCR
A new generation of technology, portable PCR, offers potential for affordable, immediate, pondside diagnosis in easy-to-use handheld kits. But will it live up to the hype?

Health & Welfare
How can Norway defeat pancreatic disease in salmon? By detecting it faster.
Polish pathogen diagnostics group will soon launch a test system that can identify the costly pancreatic disease (and six others) in just 10 minutes.
Health & Welfare
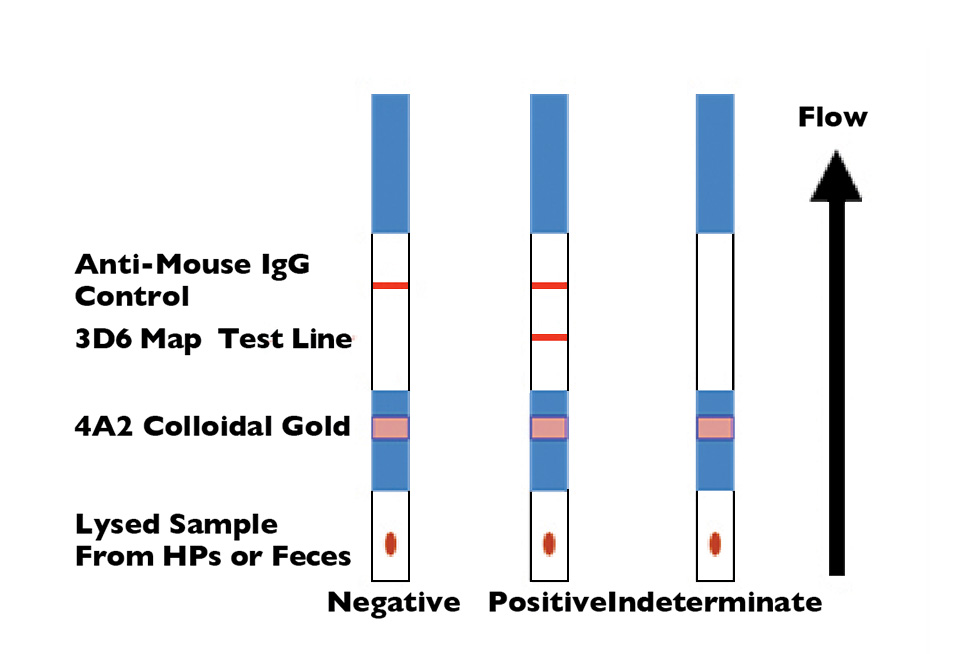
Rapid test detects NHP in penaeid shrimp
A rapid lateral-flow immunoassay using monoclonal antibody combinations can identify necrotizing hepatopancreatitis (NHP) infection in less than 15 minutes.

Health & Welfare
Challenging Pacific white postlarvae with AHPND
Study results indicate that P. vannamei challenged with AHPND in biofloc had higher survival rates than shrimp challenged in clear water.


![Ad for [membership]](https://www.globalseafood.org/wp-content/uploads/2025/07/membership_web2025_1050x125.gif)
